Connectivity analysis with PAGA - superclean
Will Macnair
Institute for Molecular Life Sciences, University of Zurich, SwitzerlandSwiss Institute of Bioinformatics (SIB), University of Zurich, SwitzerlandJanuary 05, 2022
Last updated: 2022-01-05
Checks: 5 2
Knit directory: MS_lesions/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown is untracked by Git. To know which version of the R Markdown file created these results, you’ll want to first commit it to the Git repo. If you’re still working on the analysis, you can ignore this warning. When you’re finished, you can run wflow_publish to commit the R Markdown file and build the HTML.
The global environment had objects present when the code in the R Markdown file was run. These objects can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment. Use wflow_publish or wflow_build to ensure that the code is always run in an empty environment.
The following objects were defined in the global environment when these results were created:
| Name | Class | Size |
|---|---|---|
| q | function | 1008 bytes |
The command set.seed(20210118) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version afba18d. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rprofile
Ignored: .Rproj.user/
Ignored: ._MS_lesions.sublime-project
Ignored: .log/
Ignored: MS_lesions.sublime-project
Ignored: MS_lesions.sublime-workspace
Ignored: analysis/.__site.yml
Ignored: analysis/fig_muscat_cache/
Ignored: analysis/ms00_manuscript_figures_cache/
Ignored: analysis/ms02_doublet_id_cache/
Ignored: analysis/ms03_SampleQC_cache/
Ignored: analysis/ms04_conos_cache/
Ignored: analysis/ms05_splitting_cache/
Ignored: analysis/ms06_sccaf_cache/
Ignored: analysis/ms07_soup_cache/
Ignored: analysis/ms08_modules_cache/
Ignored: analysis/ms08_modules_pseudobulk_cache/
Ignored: analysis/ms09_ancombc_cache/
Ignored: analysis/ms09_ancombc_clean_1e3_cache/
Ignored: analysis/ms09_ancombc_clean_2e3_cache/
Ignored: analysis/ms09_ancombc_mixed_cache/
Ignored: analysis/ms10_muscat_run01_cache/
Ignored: analysis/ms10_muscat_run02_cache/
Ignored: analysis/ms10_muscat_template_broad_slim_cache/
Ignored: analysis/ms10_muscat_template_fine_slim_cache/
Ignored: analysis/ms11_paga_cache/
Ignored: analysis/ms12_markers_cache/
Ignored: analysis/ms13_labelling_cache/
Ignored: analysis/ms14_lesions_cache/
Ignored: analysis/ms15_mofa_sample_gm_cache/
Ignored: analysis/ms15_mofa_sample_gm_final_meta_cache/
Ignored: analysis/ms15_mofa_sample_gm_superclean_cache/
Ignored: analysis/ms15_mofa_sample_gm_w_layers_final_meta_cache/
Ignored: analysis/ms15_mofa_sample_wm_cache/
Ignored: analysis/ms15_mofa_sample_wm_final_meta_cache/
Ignored: analysis/ms15_mofa_sample_wm_new_meta_cache/
Ignored: analysis/ms15_mofa_sample_wm_superclean_cache/
Ignored: analysis/ms15_patients_cache/
Ignored: analysis/ms15_patients_gm_cache/
Ignored: analysis/ms15_patients_sample_level_cache/
Ignored: analysis/ms15_patients_w_ms_cache/
Ignored: analysis/supp06_sccaf_cache/
Ignored: analysis/supp07_superclean_check_cache/
Ignored: analysis/supp09_ancombc_cache/
Ignored: analysis/supp09_ancombc_mixed_cache/
Ignored: analysis/supp09_ancombc_rowitch_cache/
Ignored: analysis/supp09_ancombc_superclean_cache/
Ignored: analysis/supp10_muscat_cache/
Ignored: analysis/supp10_muscat_ctrl_gm_vs_wm_cache/
Ignored: analysis/supp10_muscat_gm_layers_effects_cache/
Ignored: analysis/supp10_muscat_gsea_cache/
Ignored: analysis/supp10_muscat_heatmaps_cache/
Ignored: analysis/supp10_muscat_olg_pc1_cache/
Ignored: analysis/supp10_muscat_olg_pc2_cache/
Ignored: analysis/supp10_muscat_olg_pc_cache/
Ignored: analysis/supp10_muscat_regression_cache/
Ignored: analysis/supp10_muscat_soup_cache/
Ignored: analysis/supp10_muscat_soup_mito_cache/
Ignored: code/._ms10_muscat_fns_recover.R
Ignored: code/.recovery/
Ignored: code/jobs/._muscat_run09_2021-10-11.slurm
Ignored: code/muscat_plan.txt
Ignored: data/
Ignored: figures/
Ignored: output/
Ignored: tmp/
Untracked files:
Untracked: analysis/ms11_paga_superclean.Rmd
Untracked: analysis/supp09_ancombc_superclean.Rmd
Untracked: code/dev_dheeraj_oligo_dotplots_2021-12-08.R
Untracked: code/dev_magma_de_heatmaps_2021-12-03.R
Untracked: code/dev_plot_dotplot_sel_genes_2021-12-21.R
Untracked: code/dev_plot_fcs_selected_genes_2021-12-09.R
Untracked: code/jobs/muscat_run20_2021-12-07.slurm
Untracked: code/jobs/supp07_recalc_conos.slurm
Untracked: code/supp07_recalc_conos.R
Unstaged changes:
Modified: analysis/fig_muscat.Rmd
Modified: analysis/ms00_manuscript_figures.Rmd
Modified: analysis/ms11_paga.Rmd
Modified: analysis/ms13_labelling.Rmd
Modified: analysis/ms14_lesions.Rmd
Modified: analysis/ms15_mofa_sample_wm_superclean.Rmd
Modified: analysis/supp07_superclean_check.Rmd
Modified: code/dev_de_w_contamation_2021-10-25.R
Modified: code/dev_edger_on_mofa_20210804.R
Modified: code/fig_muscat.R
Modified: code/ms04_conos.R
Modified: code/ms11_paga.R
Modified: code/ms11_paga_fns.py
Modified: code/ms14_lesions.R
Modified: code/supp07_superclean.R
Modified: code/supp10_muscat.R
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
There are no past versions. Publish this analysis with wflow_publish() to start tracking its development.
Setup / definitions
Libraries
Helper functions
source('code/ms00_utils.R')
source('code/ms09_ancombc_mixed.R')
source('code/ms11_paga.R')
use_condaenv('scanpy', required=TRUE)
source_python('code/ms11_paga_fns.py')
source('code/ms03_SampleQC.R')Inputs
# define inputs
qc_dir = 'output/ms03_SampleQC'
qc_f = 'output/ms03_SampleQC/ms_qc_dt.txt'
graph_f = 'output/ms04_conos/conos_graph_2021-02-11.txt'
# define run
labels_f = 'data/byhand_markers/validation_markers_2021-05-31.csv'
labelled_f = 'output/ms13_labelling/conos_labelled_2021-05-31.txt.gz'
meta_f = "data/metadata/metadata_checked_assumptions_2021-10-08.xlsx"Outputs
save_dir = 'output/ms11_paga'
if (!dir.exists(save_dir))
dir.create(save_dir)
date_tag = '2022-01-04_clean'
seed = 20220104
# do in parallel
n_cores = 16
# identifying strange samples
neuro_mad_cut = 2
log_n_mad_cut = 3
min_counts = 2000
min_feats = 500
max_mito = 0.1
max_splice = 2
min_cells = 500
# where to save files with subgraphs?
subg_pat = sprintf('%s/subgraph_%s_%s.txt.gz', save_dir, '%s', date_tag)
# define subsets to run
subset_list = list(
all = list(),
healthy_WM = list(lesion_type = c('WM')),
MS_WM = list(lesion_type = c('NAWM', 'AL', 'CAL', 'CIL', 'RL')),
healthy_GM = list(lesion_type = c('GM')),
MS_GM = list(lesion_type = c('NAGM', 'GML'))
)
lesion_list = lapply(lesion_ord, function(l)
list(lesion_type = l)) %>%
setNames(paste0('lesion_', lesion_ord)
)
xy_ref_list = c(rep('WM', 6), rep('GM', 3)) %>%
paste0('healthy_', .) %>% setNames(names(lesion_list))
# what to look at when comparing edges across samples?
sel_broad = c('Oligodendrocytes', 'OPCs / COPs')
assert_that(all(sel_broad %in% broad_ord))[1] TRUE# define ancom outputs to use
ancom_dir = 'output/ms09_ancombc'
ancom_stamp = '2021-11-12'
ancom_tags = c(WM = "lesions_WM", GM = "lesions_GM_4pcs")
ancom_fs = sapply(ancom_tags, function(t)
sprintf('%s/ancombc_bootstrap_%s_%s.txt.gz', ancom_dir, t, ancom_stamp)) %>%
setNames(names(ancom_tags))Load inputs
qc_dt = get_qc_dt(qc_dir, qc_f)
qc_keep = qc_dt[
(log_counts >= log10(min_counts)) &
(log_feats >= log10(min_feats)) &
(logit_mito <= qlogis(max_mito)) &
(splice_ratio <= max_splice)
] %>%
.[, N_sample := .N, by = sample_id] %>%
.[ N_sample >= min_cells ]meta_dt = load_meta_dt_from_xls(meta_f)
labels_dt = load_names_dt(labels_f) %>%
.[, cluster_id := type_fine]
conos_dt = load_labelled_dt(labelled_f, labels_f) %>%
merge(meta_dt, by = 'sample_id') %>%
add_neuro_props(mad_cut = neuro_mad_cut)# add lib size cutoff
conos_dt = conos_dt %>%
merge(qc_keep, by = c("sample_id", "cell_id"))# # check for any outliers
# size_chks = calc_size_outliers(conos_dt, mad_cut = log_n_mad_cut)
# message("these samples excluded to outlier sample sizes:")
# print(size_chks[ size_ok == FALSE ])
# # exclude them from conos
# ok_samples = size_chks[ size_ok == TRUE ]$sample_id
# conos_dt = conos_dt[ (sample_id %in% ok_samples) & (neuro_ok == TRUE) ]
conos_dt = conos_dt[ (neuro_ok == TRUE) ]# make list of all samples
samples = unique(conos_dt$sample_id)
sample_list = lapply(samples, function(s) list(sample_id = s)) %>%
setNames(paste0('sample_', samples))
conos_tidy = conos_dt[, .(sample_id, matter, lesion_type, cell_id, type_broad, type_fine)]ancom_ls = ancom_fs %>% lapply(function(f) fread(f) %>%
.calc_boots_sum(min_effect = 0.1, q_cut = 0.05, signif_cut = 0.8)) %>%
setNames(names(ancom_fs))Processing / calculations
# make subgraphs for each specification
set.seed(seed)
calc_subgraphs(conos_tidy, subset_list, graph_f, subg_pat, n_sample = 1e5)already done!NULLcalc_subgraphs(conos_tidy, lesion_list, graph_f, subg_pat, n_sample = 1e5)already done!NULLcalc_subgraphs(conos_tidy, sample_list, graph_f, subg_pat, n_sample = 0)already done!NULL# run on each sample
if (!check_all_done(sample_list, save_dir, date_tag)) {
bpparam = MulticoreParam(workers = n_cores)
bpstop()
bpstart()
ignore_me = bplapply(seq_along(sample_list), function(i) {
spec_n = names(sample_list)[[i]]
subg_f = sprintf(subg_pat, spec_n)
run_paga(save_dir, spec_n, date_tag, r_to_py(as.data.frame(conos_tidy)),
subg_f, group_var = 'type_fine')
}, BPPARAM = bpparam)
bpstop()
}
# run on each lesion type
if (!check_all_done(lesion_list, save_dir, date_tag)) {
n_lesions = length(lesion_list)
bpparam = MulticoreParam(workers = min(n_lesions, n_cores))
bpstop()
bpstart()
bplapply(seq_along(lesion_list), function(i) {
spec_n = names(lesion_list)[[i]]
subg_f = sprintf(subg_pat, spec_n)
run_paga(save_dir, spec_n, date_tag, r_to_py(as.data.frame(conos_tidy)),
subg_f, group_var = 'type_fine')
}, BPPARAM = bpparam)
bpstop()
}
# run on each subset
if (!check_all_done(subset_list, save_dir, date_tag)) {
bpparam = MulticoreParam(workers = length(subset_list))
bpstop()
bpstart()
bplapply(seq_along(subset_list), function(i) {
# define things
spec_n = names(subset_list)[[i]]
subg_f = sprintf(subg_pat, spec_n)
# run paga
run_paga(save_dir, spec_n, date_tag, r_to_py(as.data.frame(conos_tidy)),
subg_f, group_var = 'type_fine')
}, BPPARAM = bpparam)
bpstop()
}edges_all = calc_edges_over_samples(sample_list, date_tag,
sel_broad, labels_dt)
edges_all = merge(edges_all, meta_dt, by = 'sample_id')Analysis
PAGA on all celltypes
Split by GM / WM + condition
for (nn in names(subset_list)) {
cat('#### ', nn, '\n')
suppressWarnings({print(plot_paga_outputs(save_dir, nn, date_tag, 'all',
conos_dt, subset_list[[nn]], labels_dt, xy_ref_n = nn))})
cat('\n\n')
}all
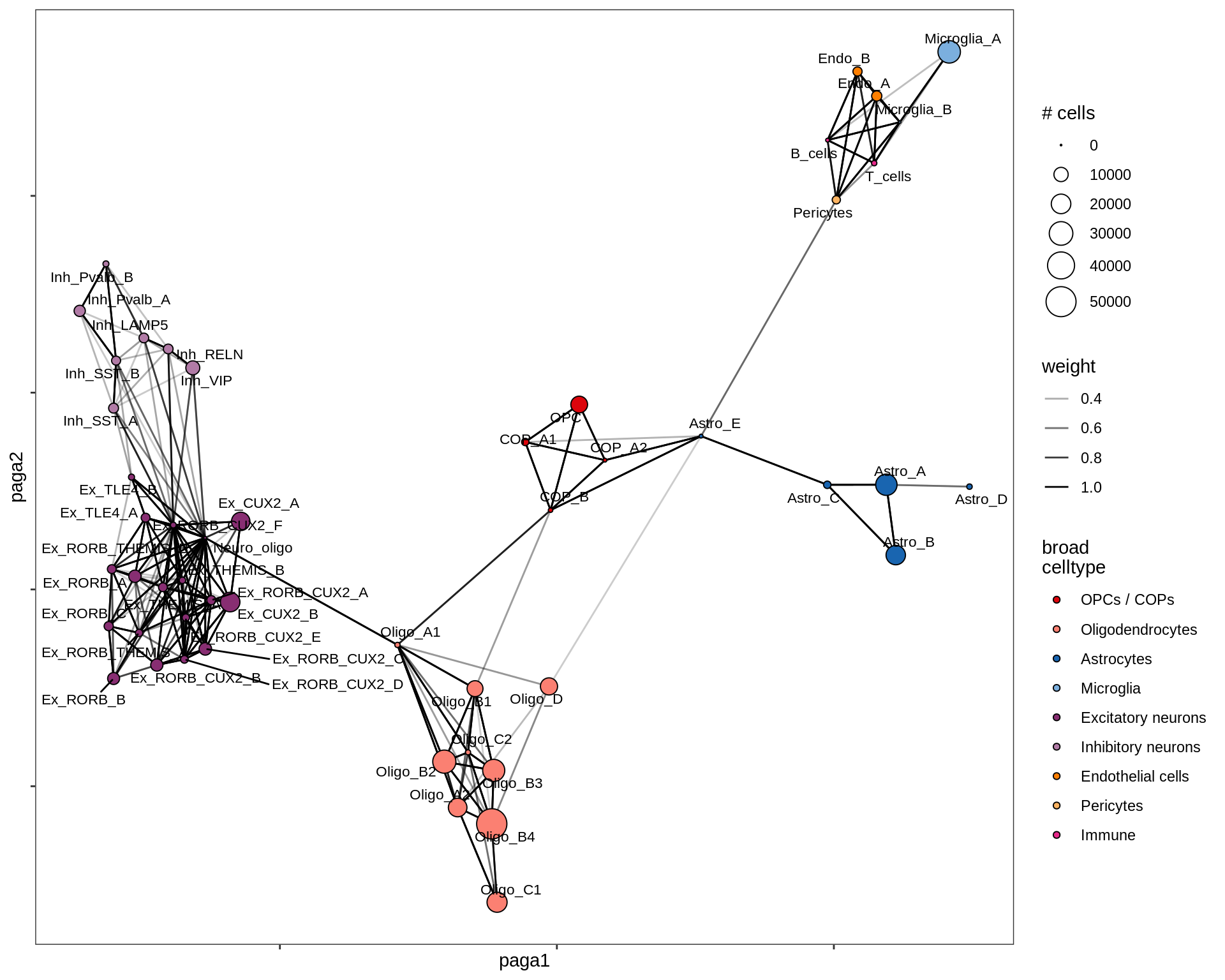
healthy_WM
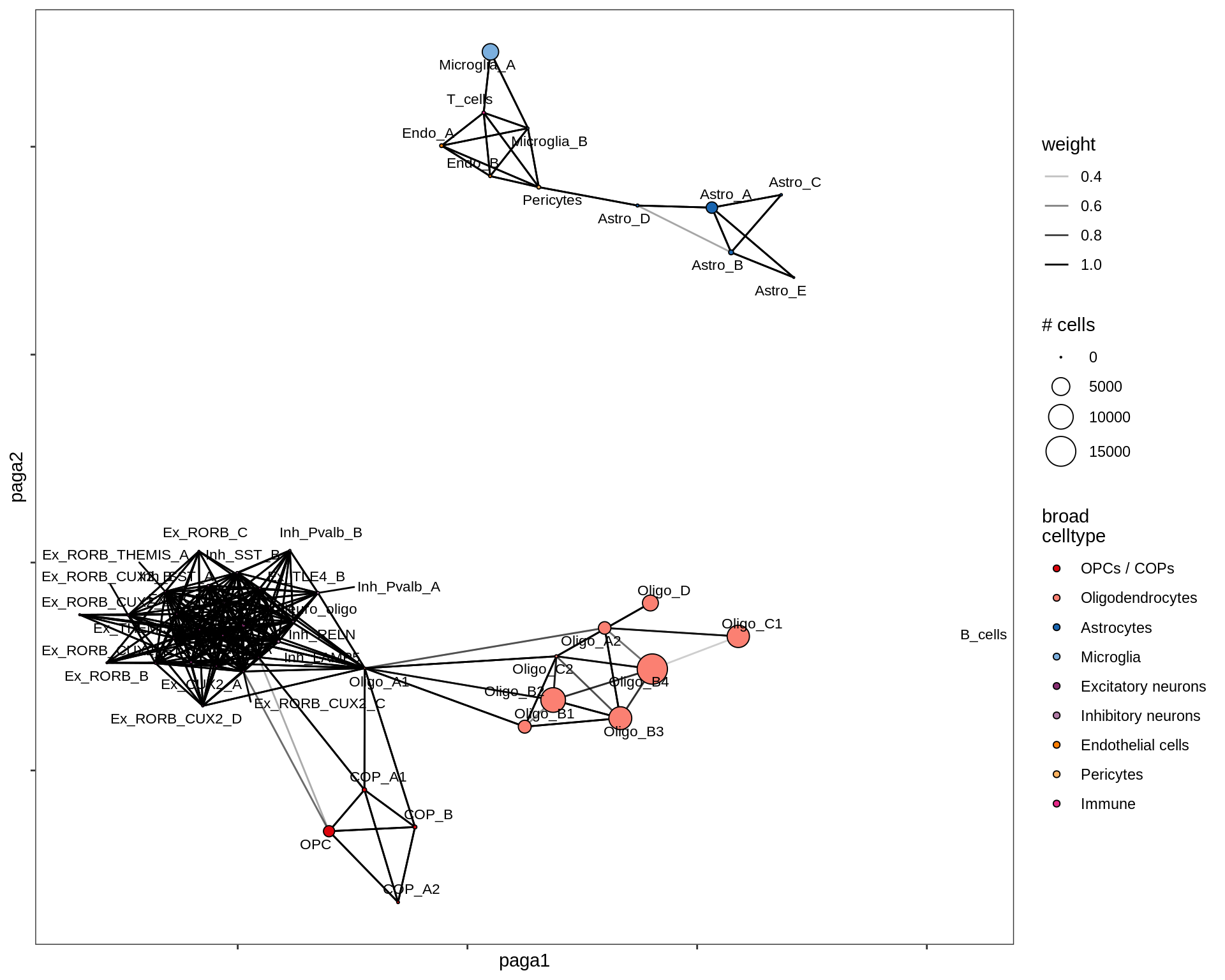
MS_WM
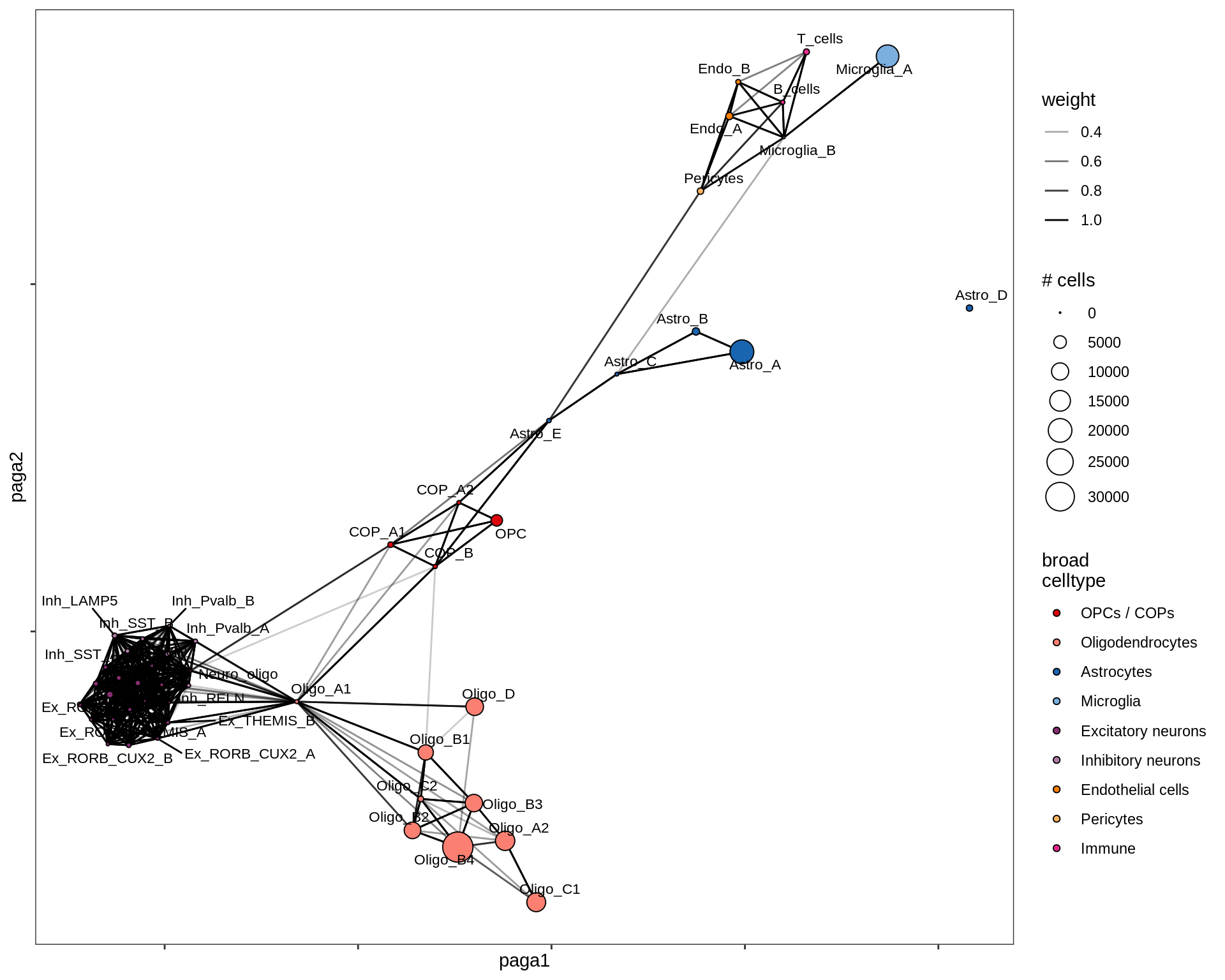
healthy_GM
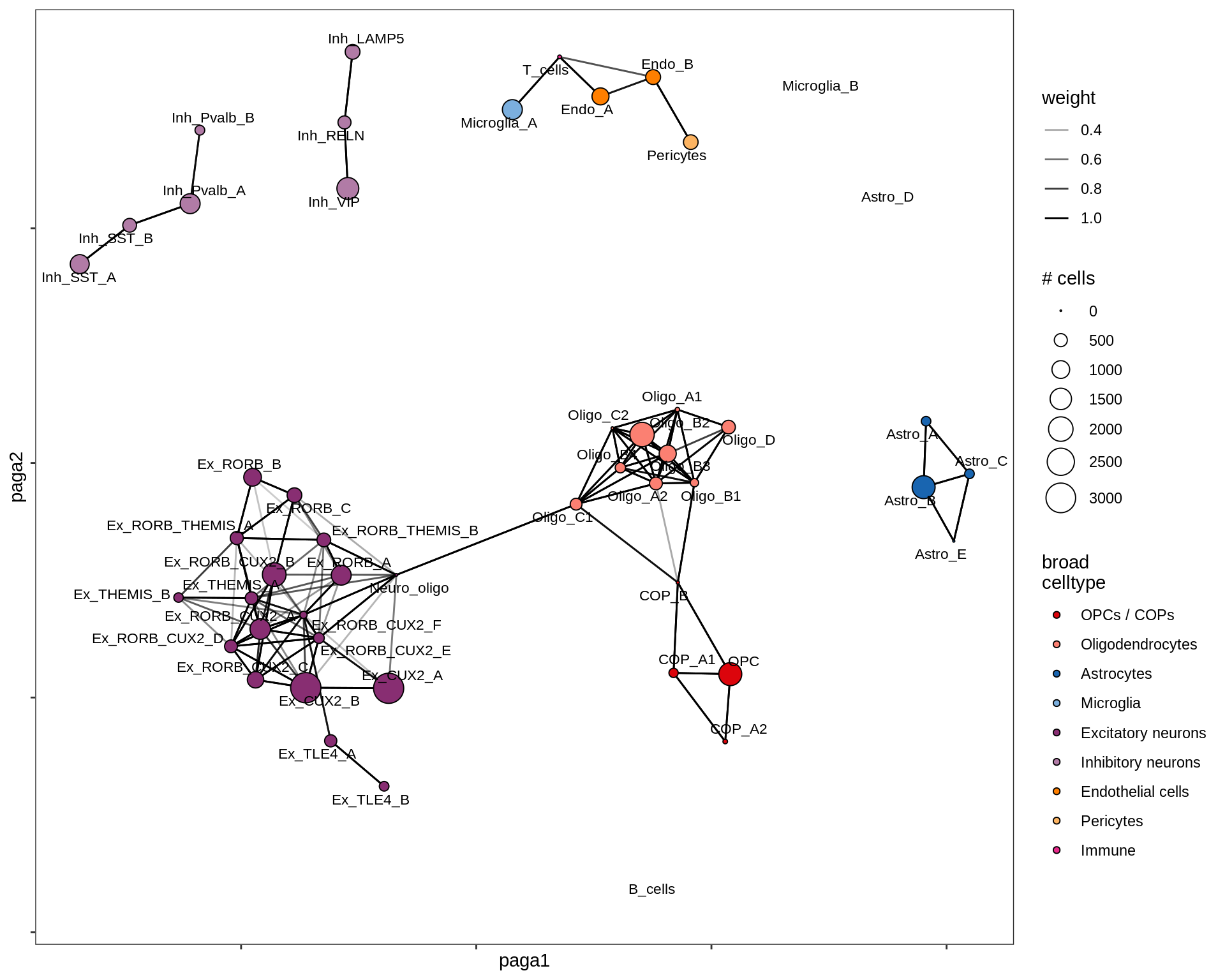
MS_GM
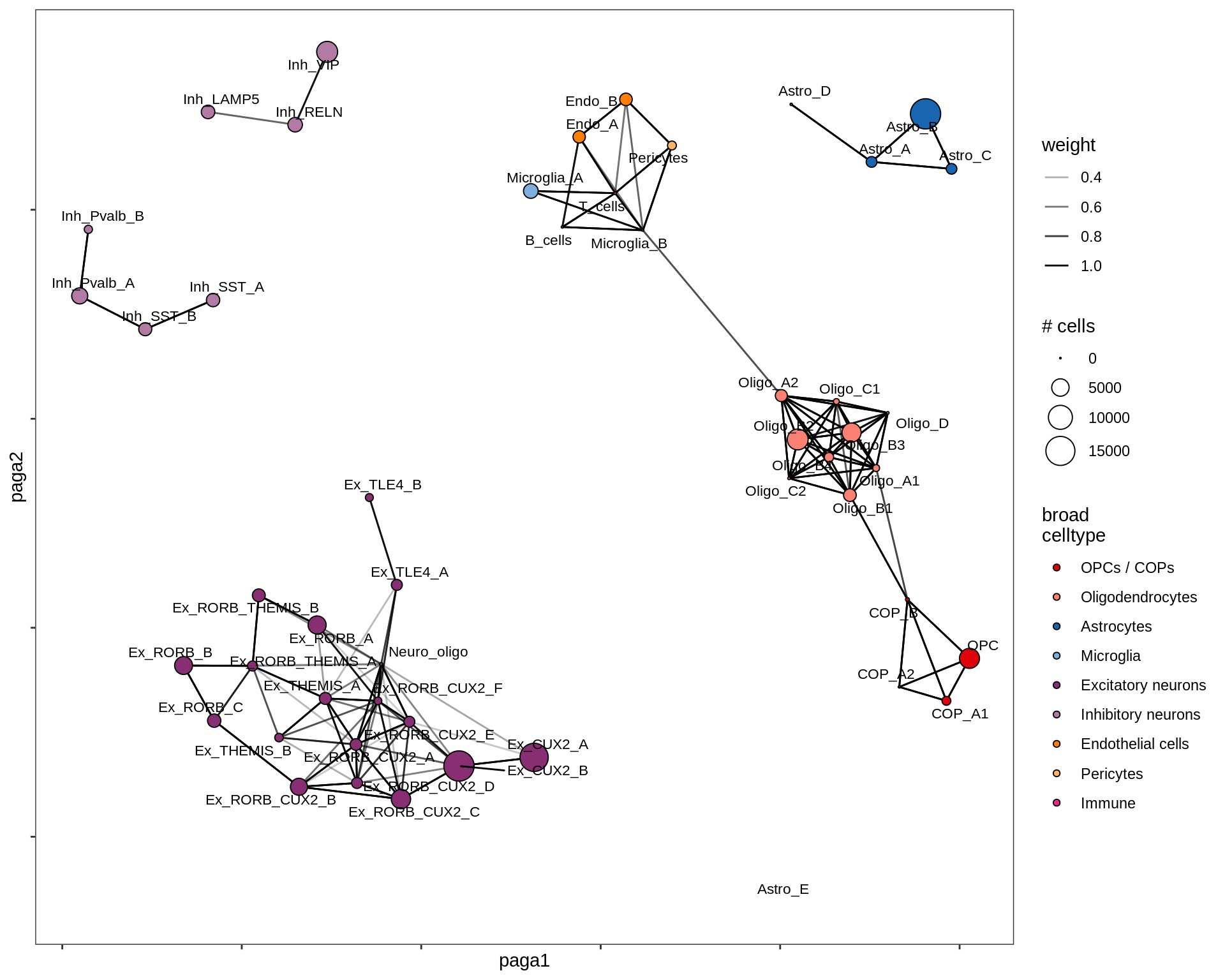
Split by lesion type
for (nn in names(lesion_list)) {
cat('#### ', nn, '\n')
suppressWarnings({print(plot_paga_outputs(save_dir, nn, date_tag, 'all',
conos_dt, lesion_list[[nn]], labels_dt, xy_ref_n = 'all'))})
cat('\n\n')
}lesion_WM
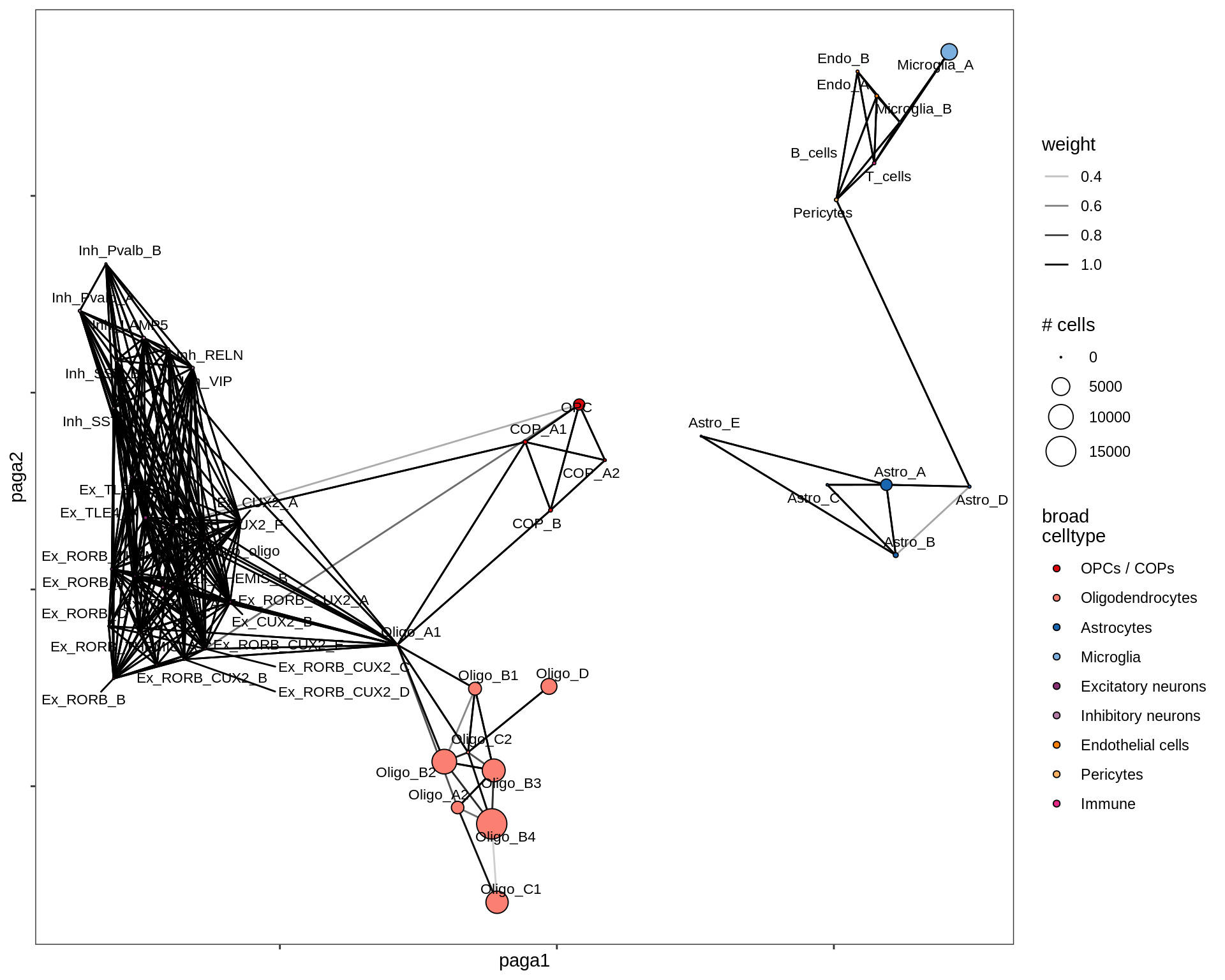
lesion_NAWM
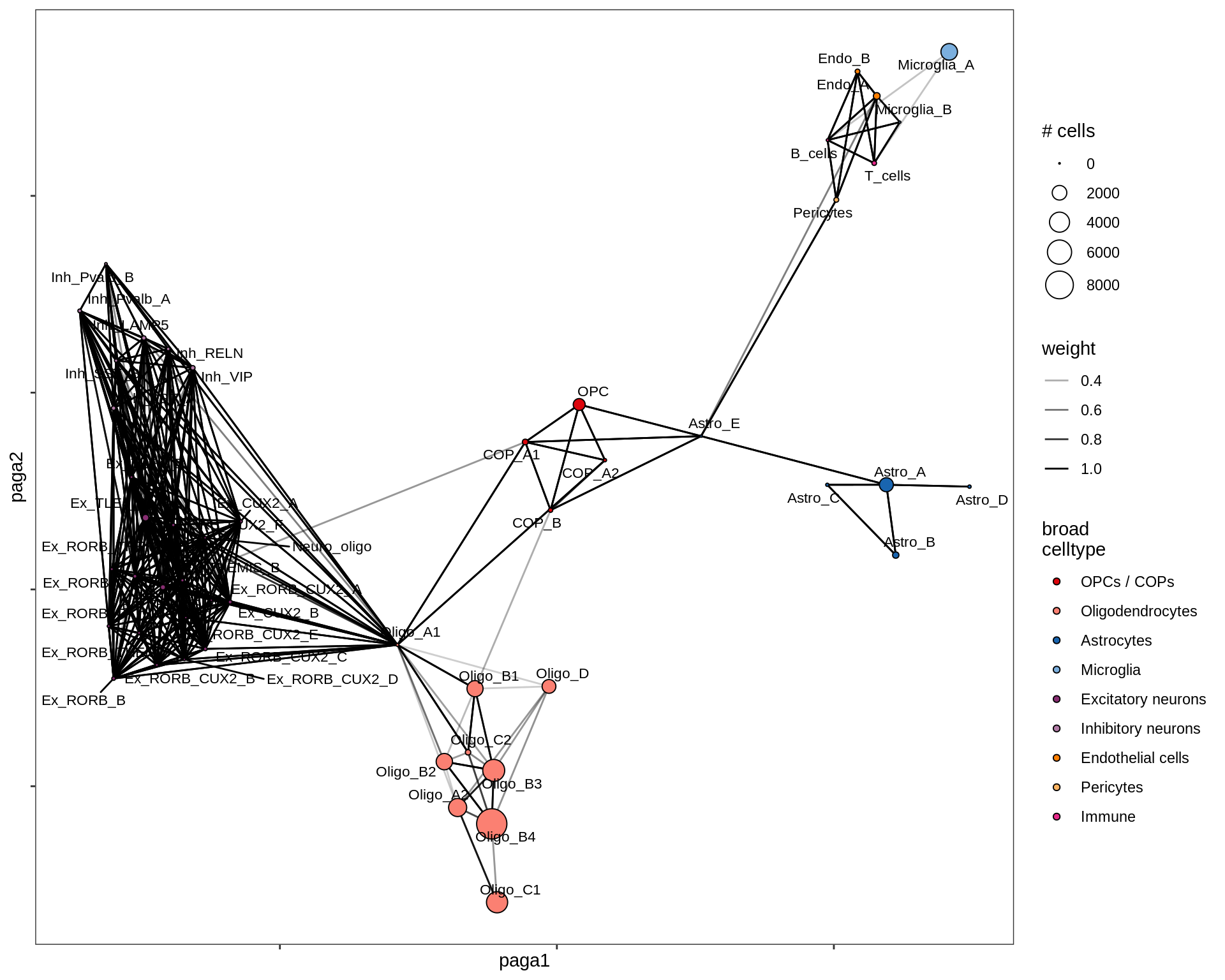
lesion_AL
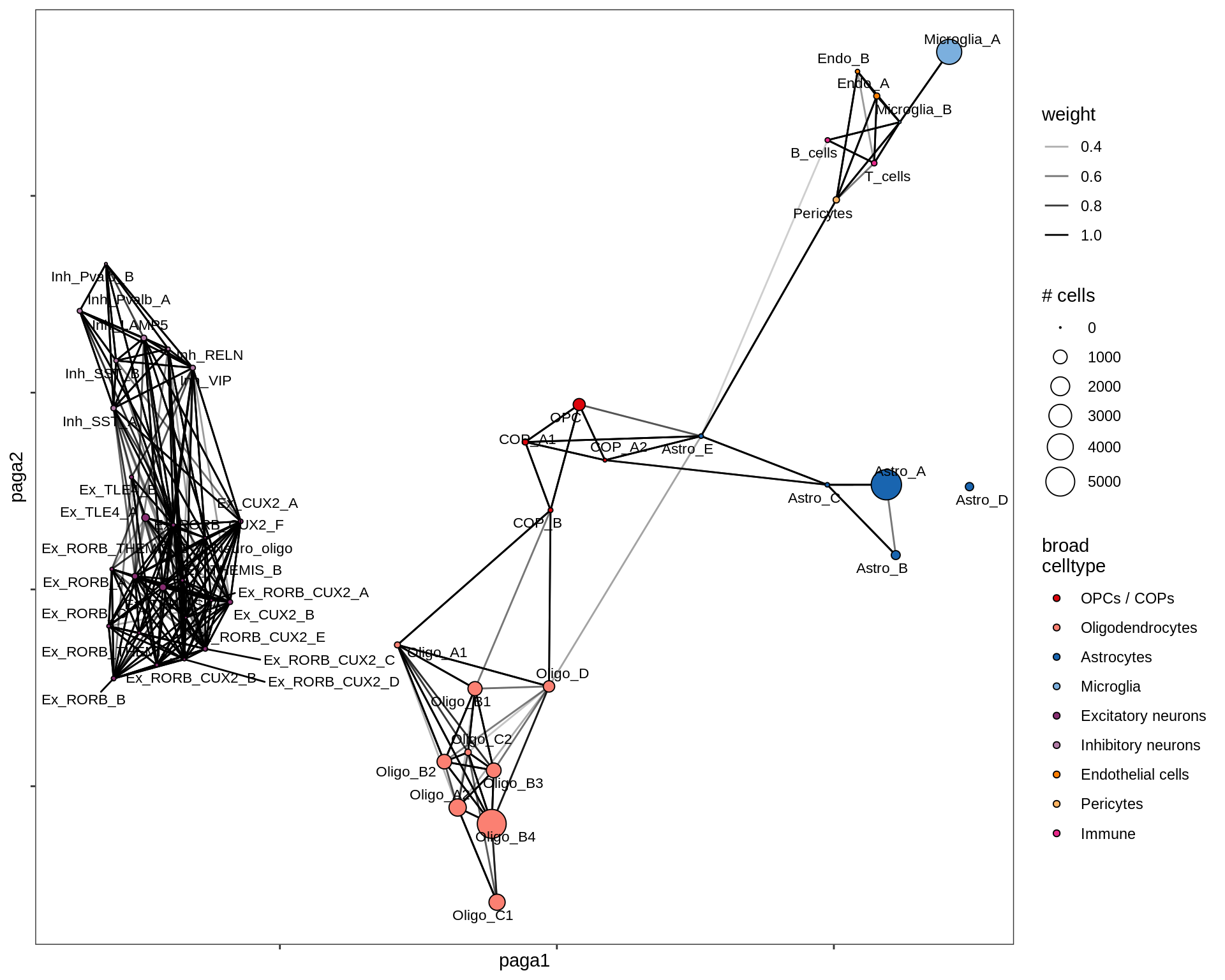
lesion_CAL
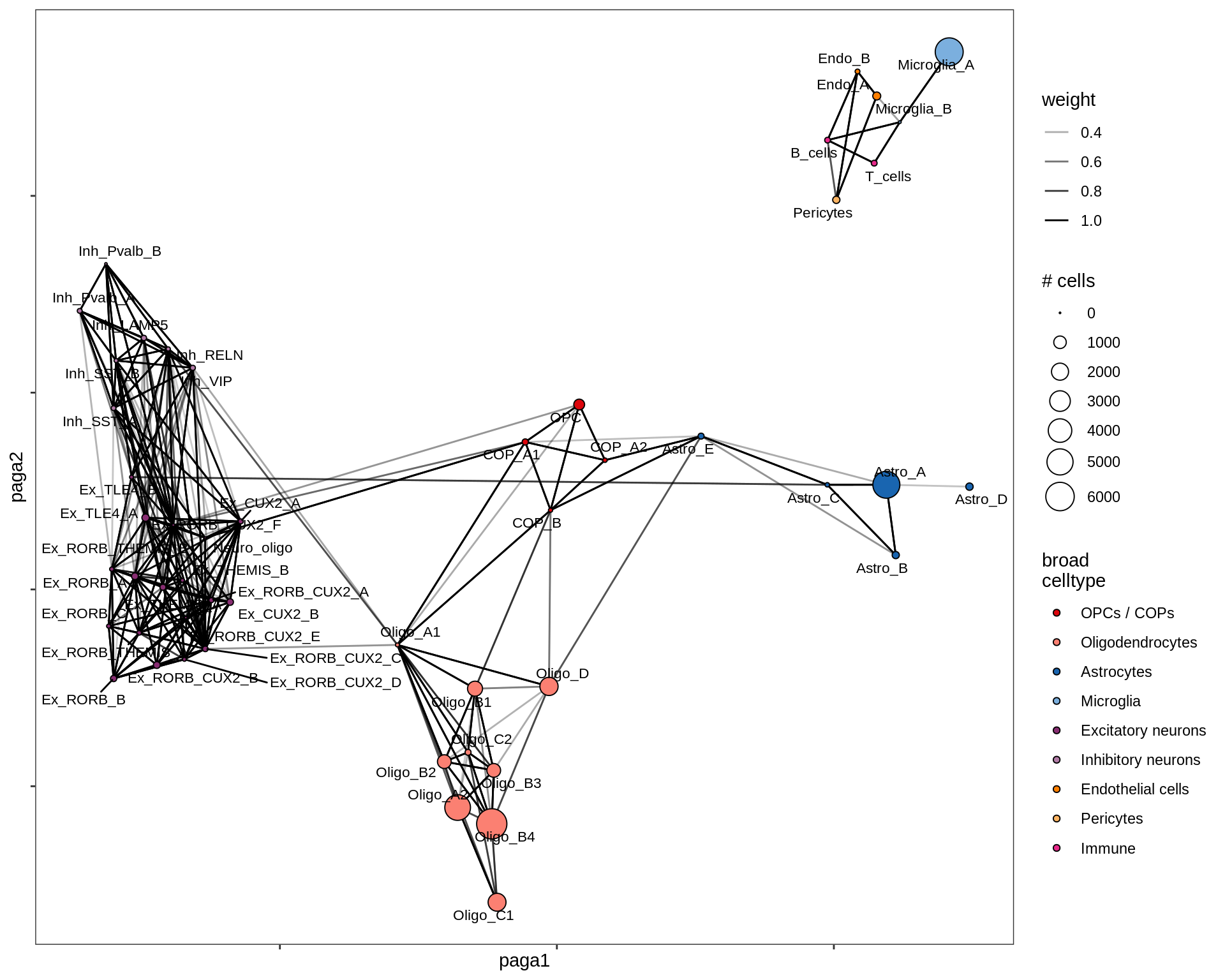
lesion_CIL
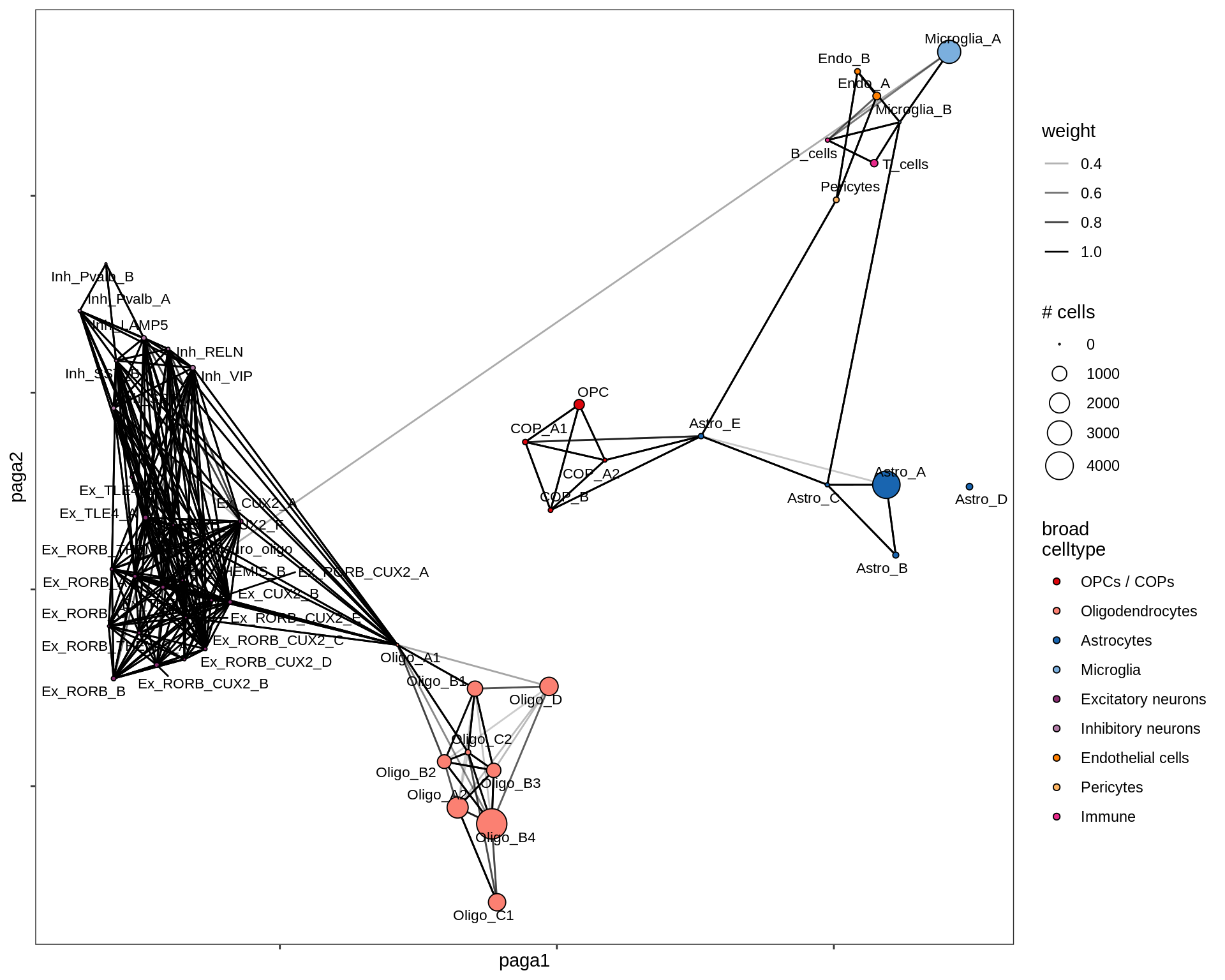
lesion_RL
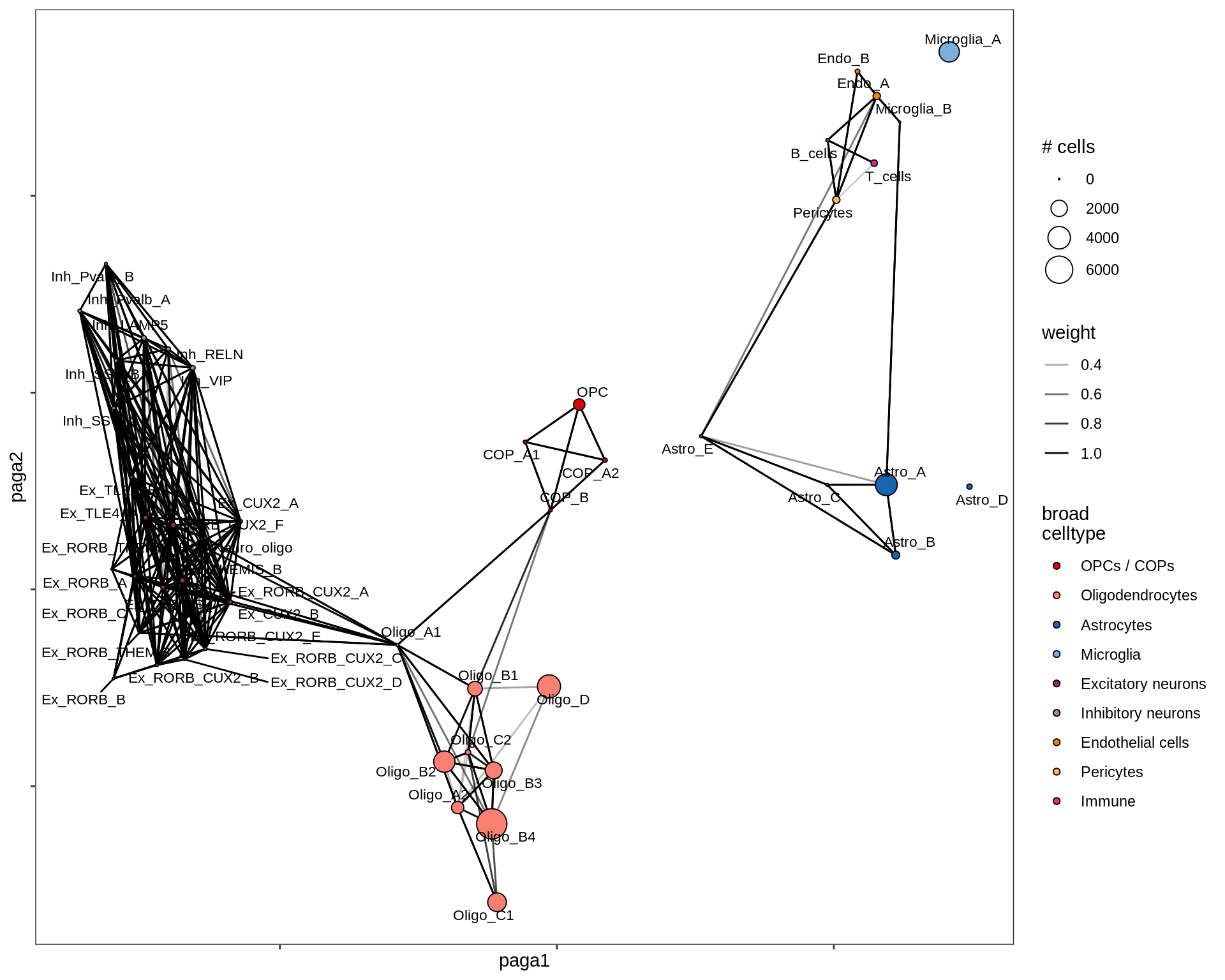
lesion_GM
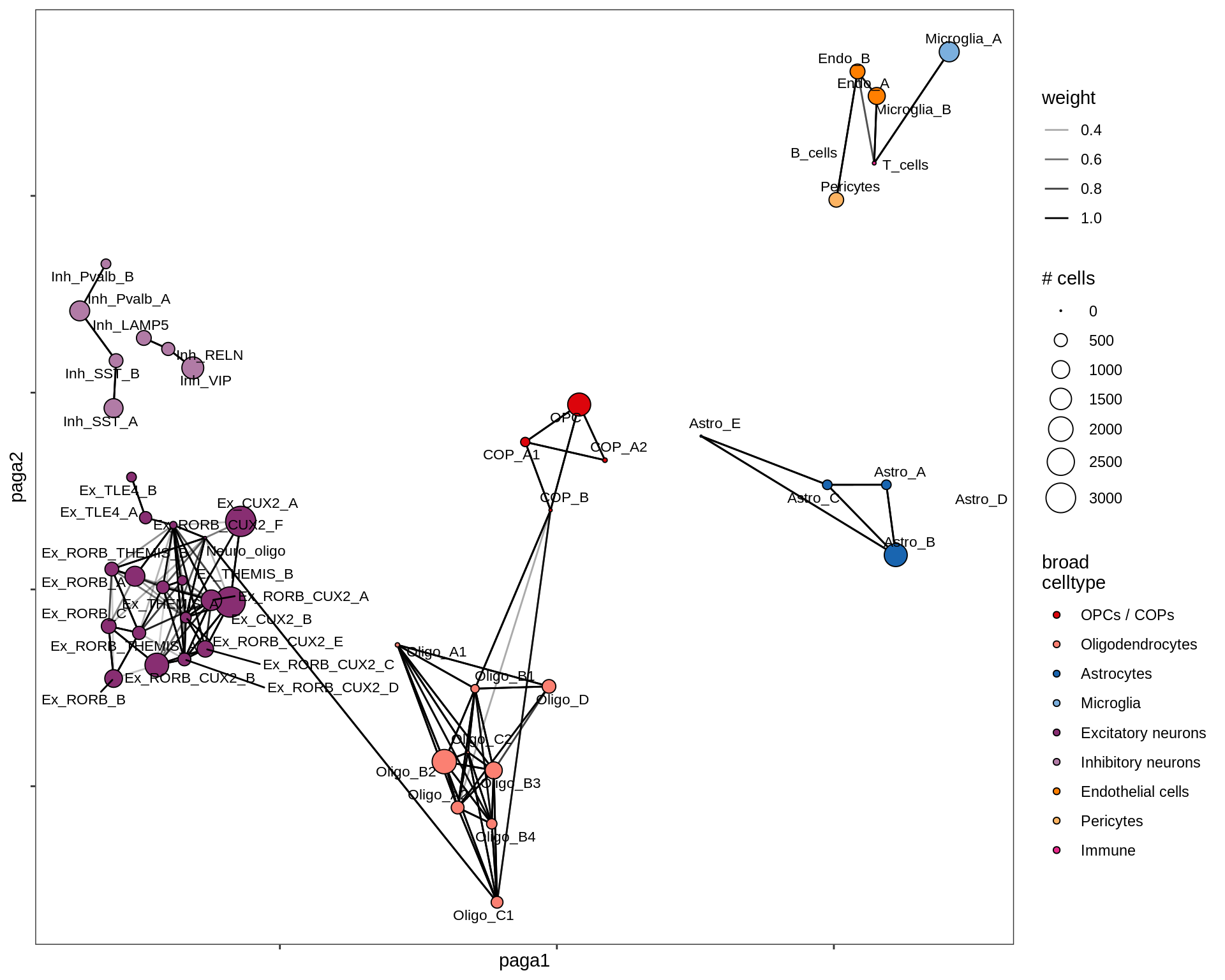
lesion_NAGM
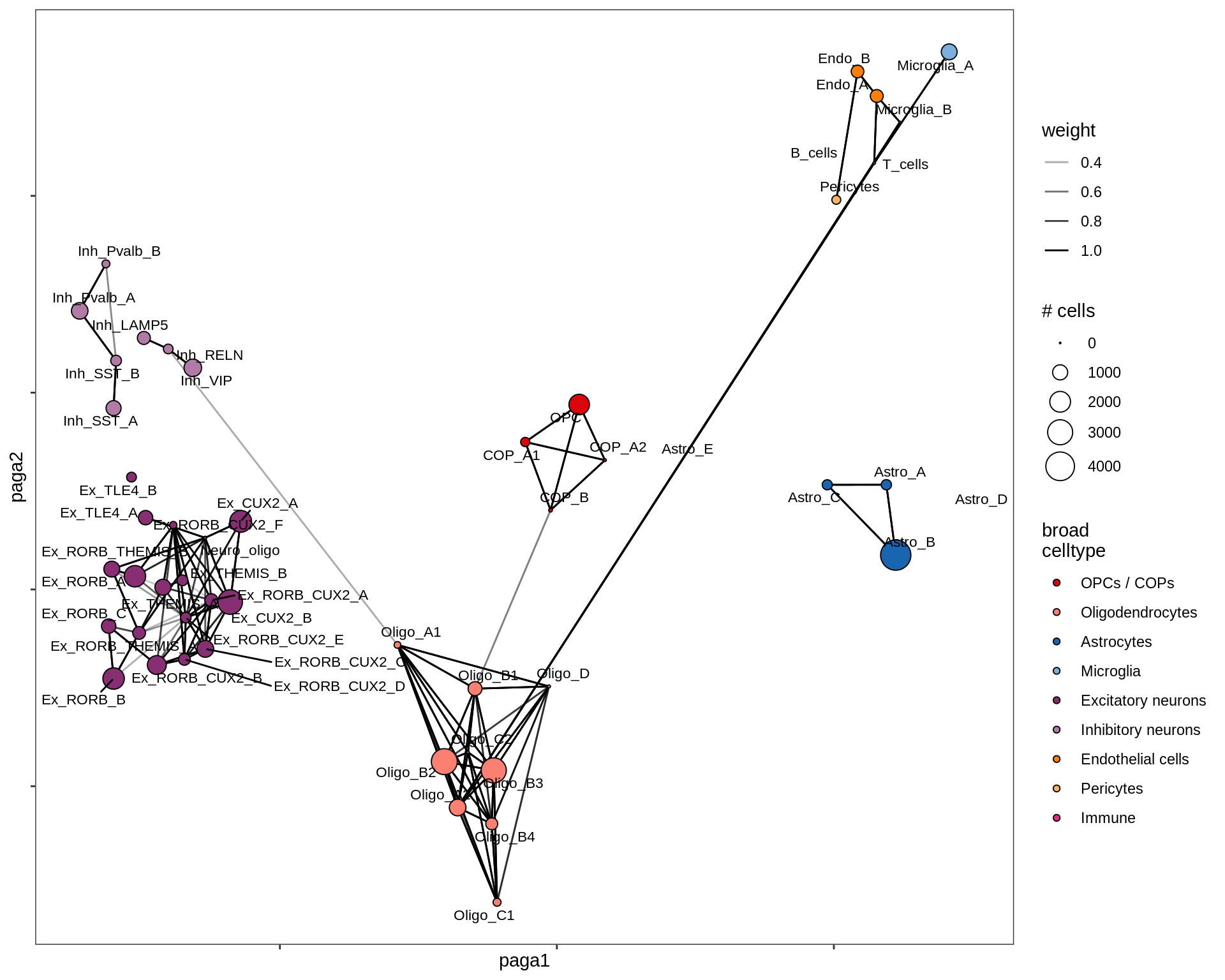
lesion_GML
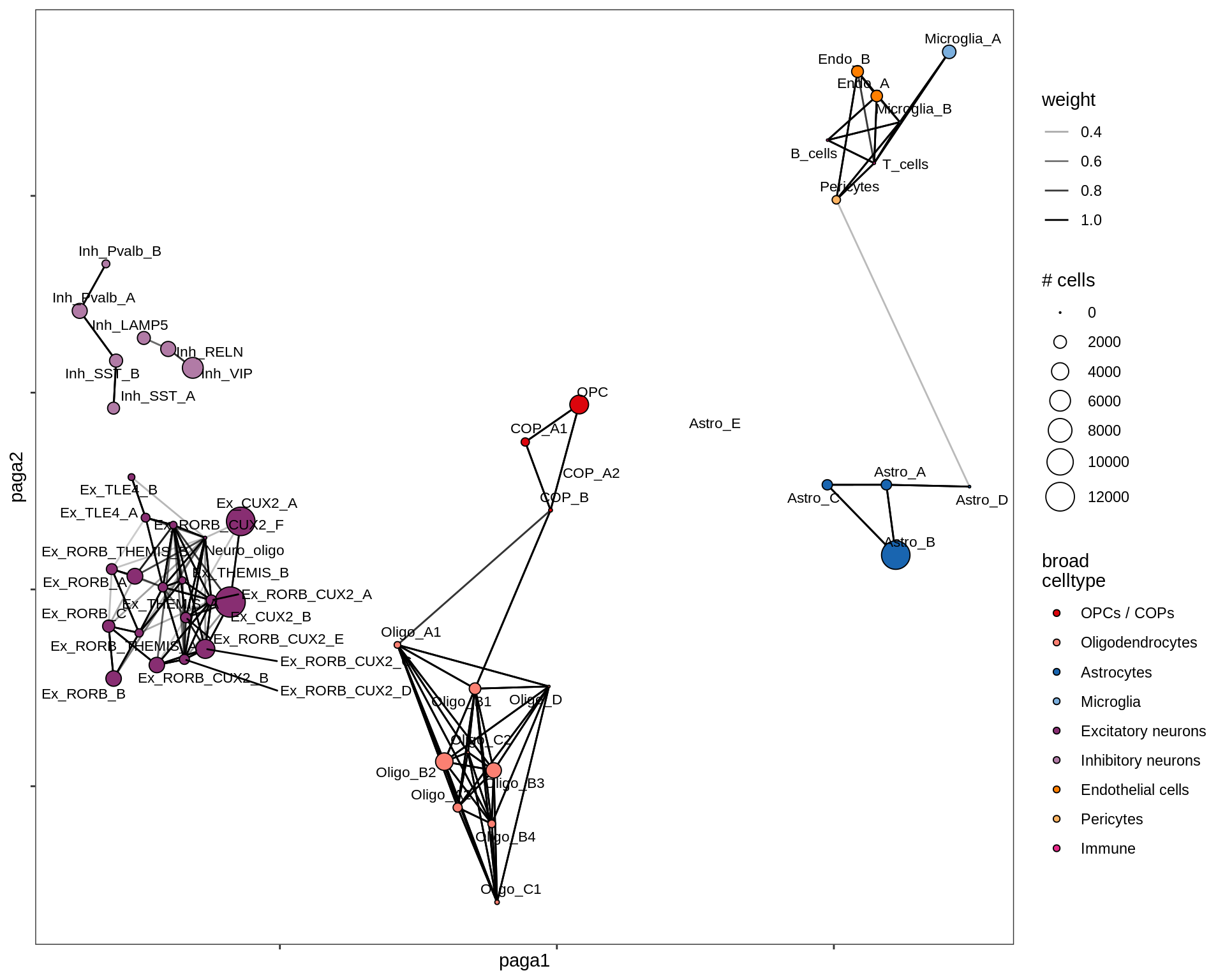
PAGA on oligo + OPC compartment
Split by GM / WM + condition
for (nn in names(subset_list)) {
cat('#### ', nn, '\n')
suppressWarnings({print(plot_paga_outputs(save_dir, nn, date_tag, 'olg',
conos_dt, subset_list[[nn]], labels_dt, xy_ref_n = nn))})
cat('\n\n')
}all
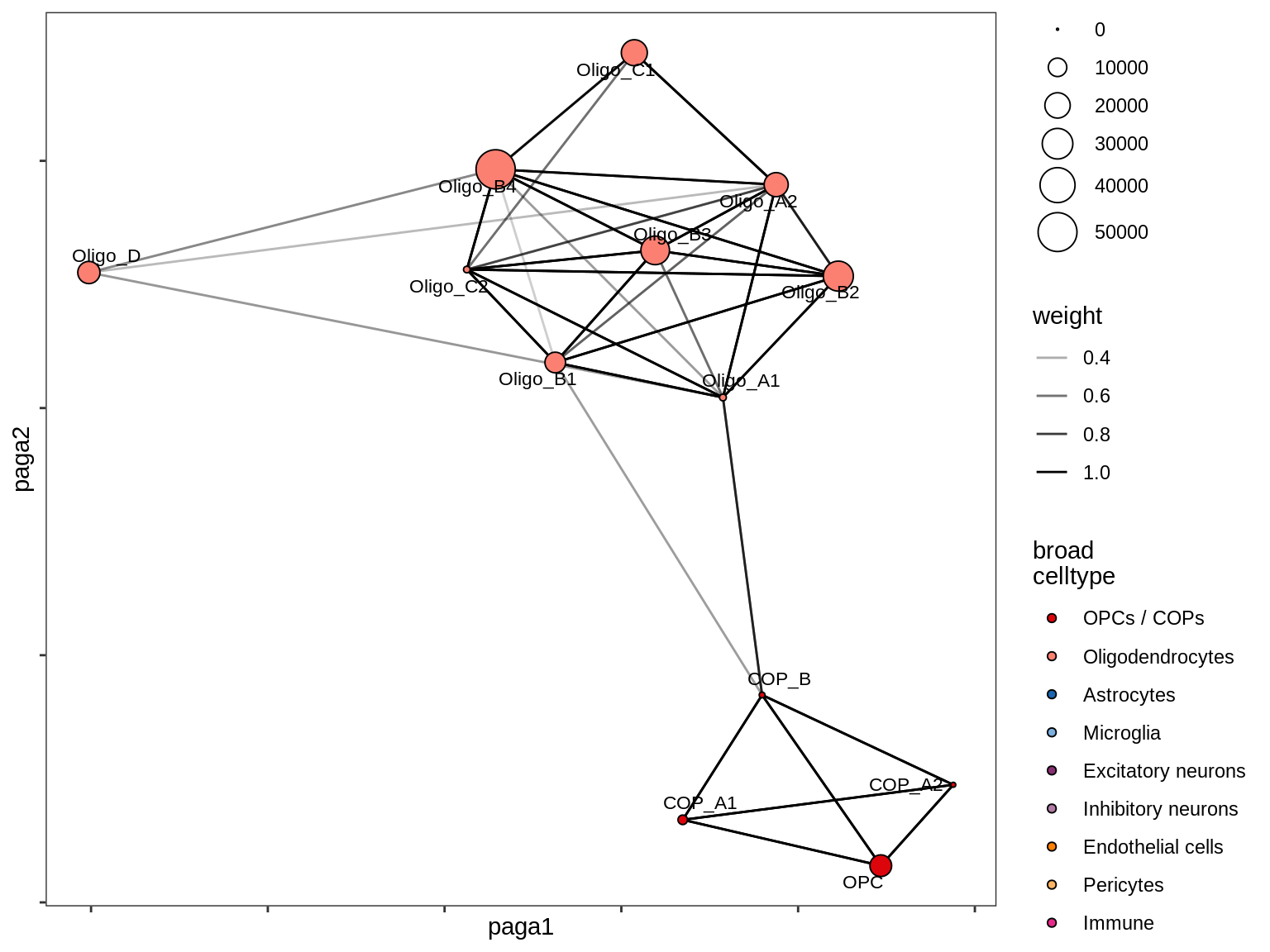
healthy_WM
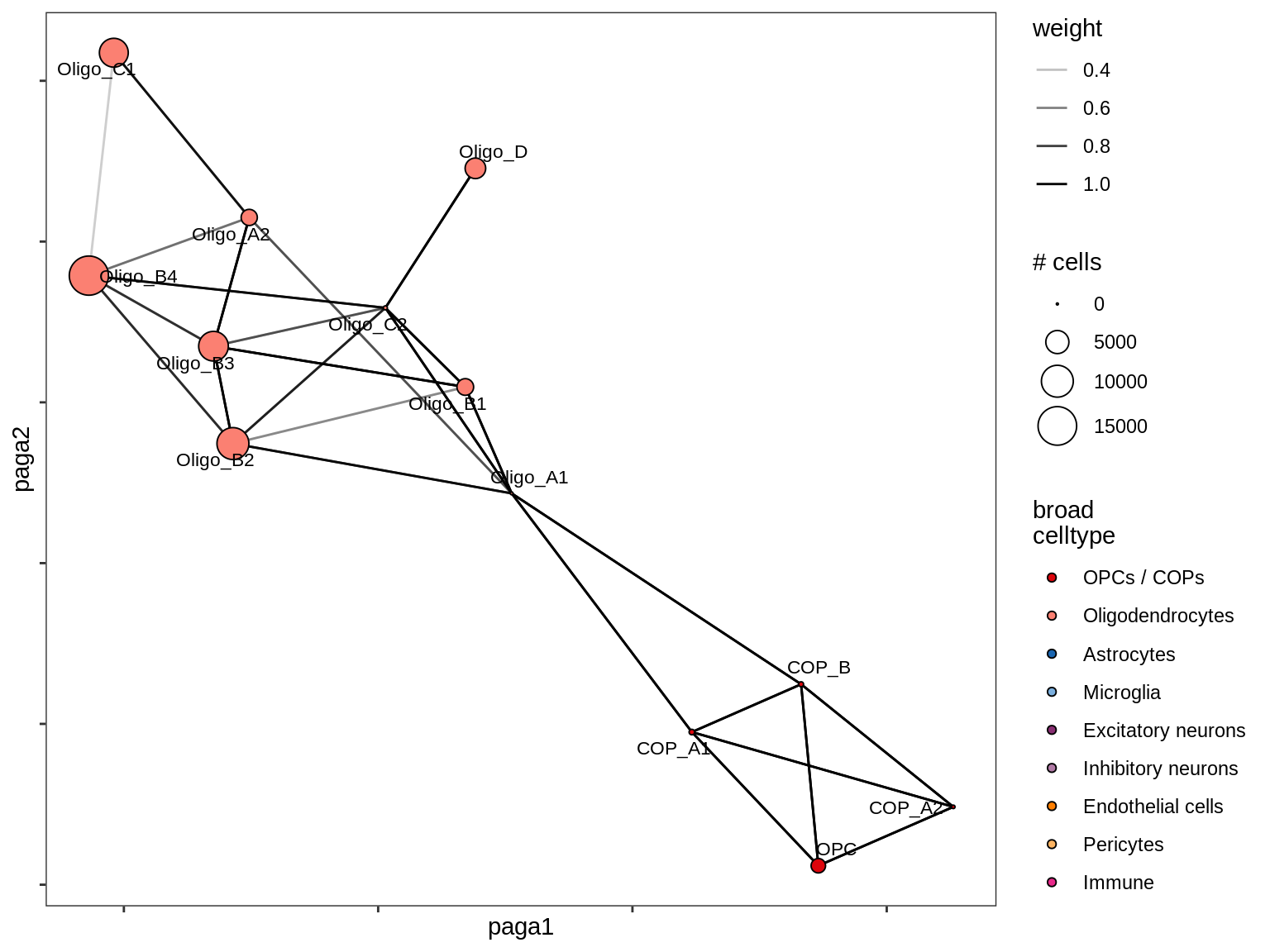
MS_WM
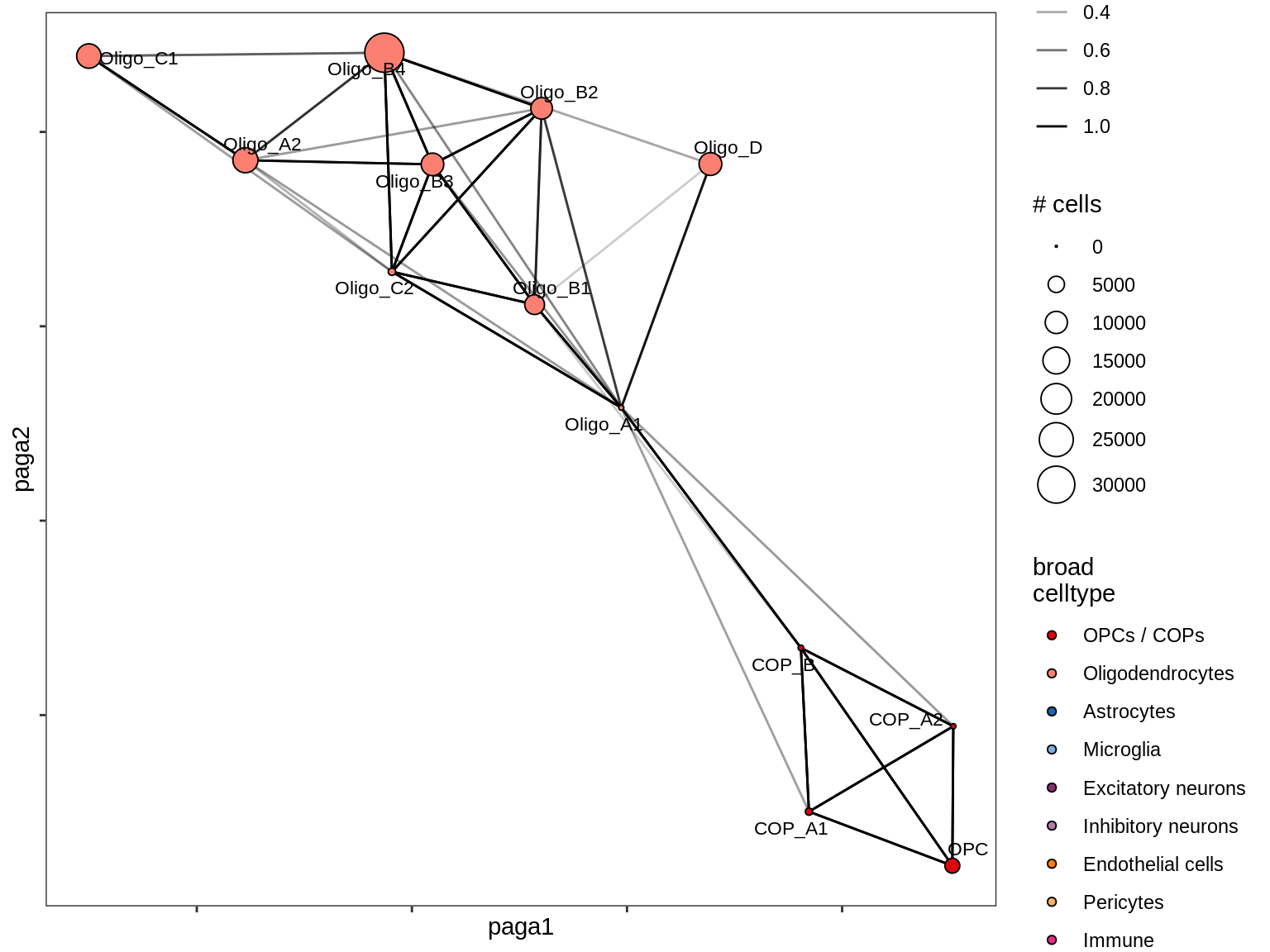
healthy_GM
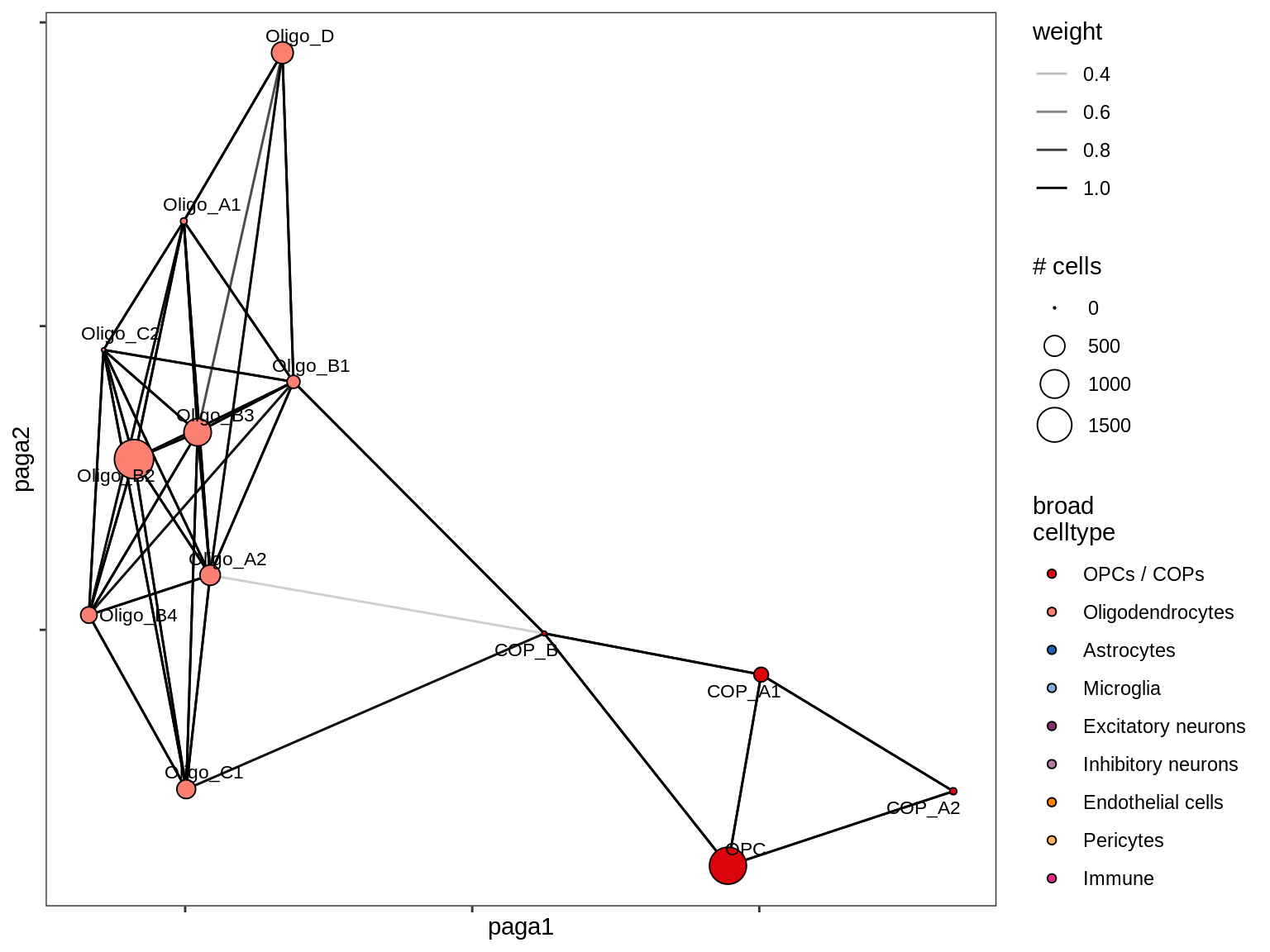
MS_GM
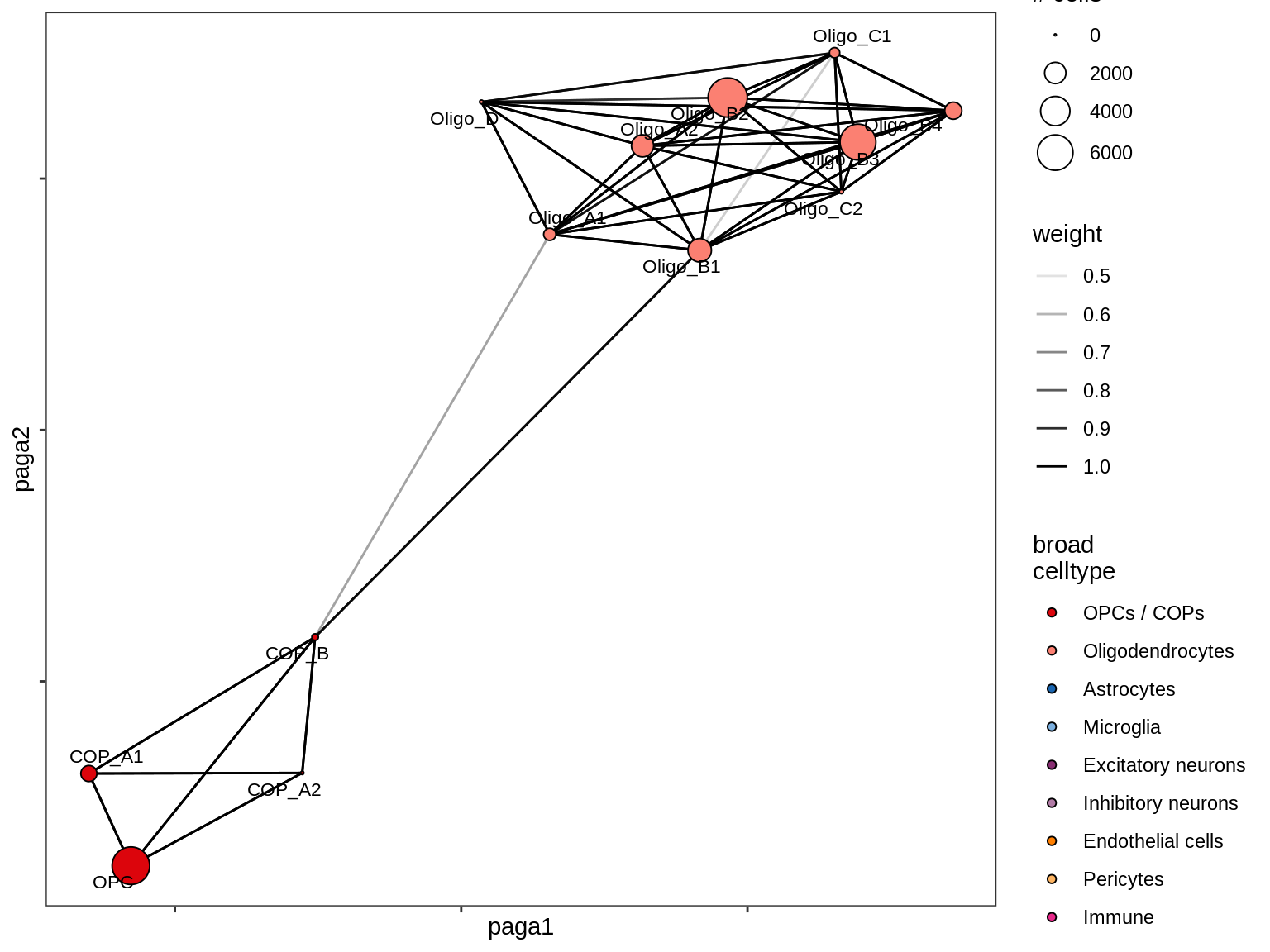
Split by lesion type
for (nn in names(lesion_list)) {
cat('#### ', nn, '\n')
suppressWarnings({print(plot_paga_outputs(save_dir, nn, date_tag, 'olg',
conos_dt, lesion_list[[nn]], labels_dt, xy_ref_n = 'all'))})
cat('\n\n')
}lesion_WM
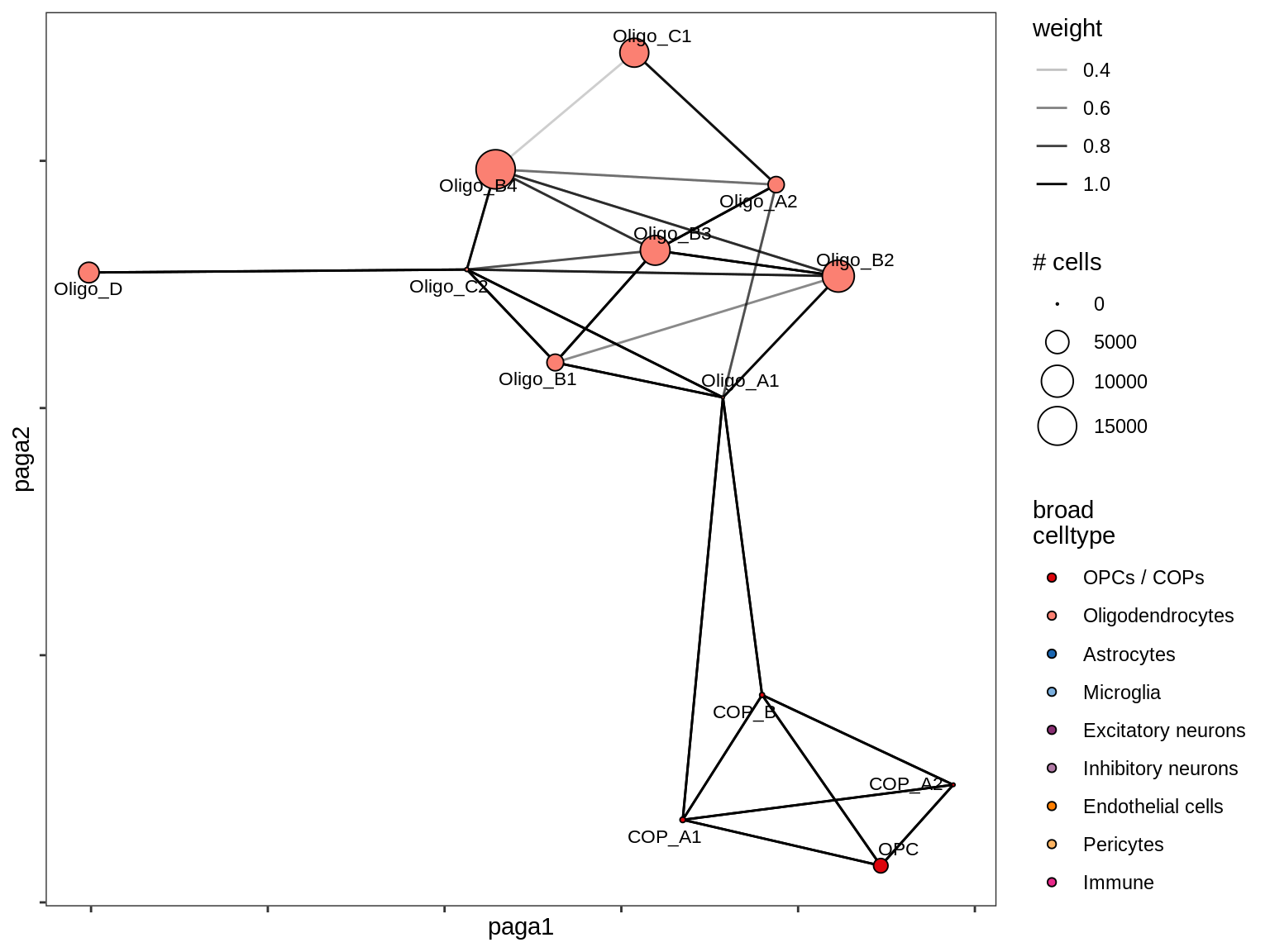
lesion_NAWM
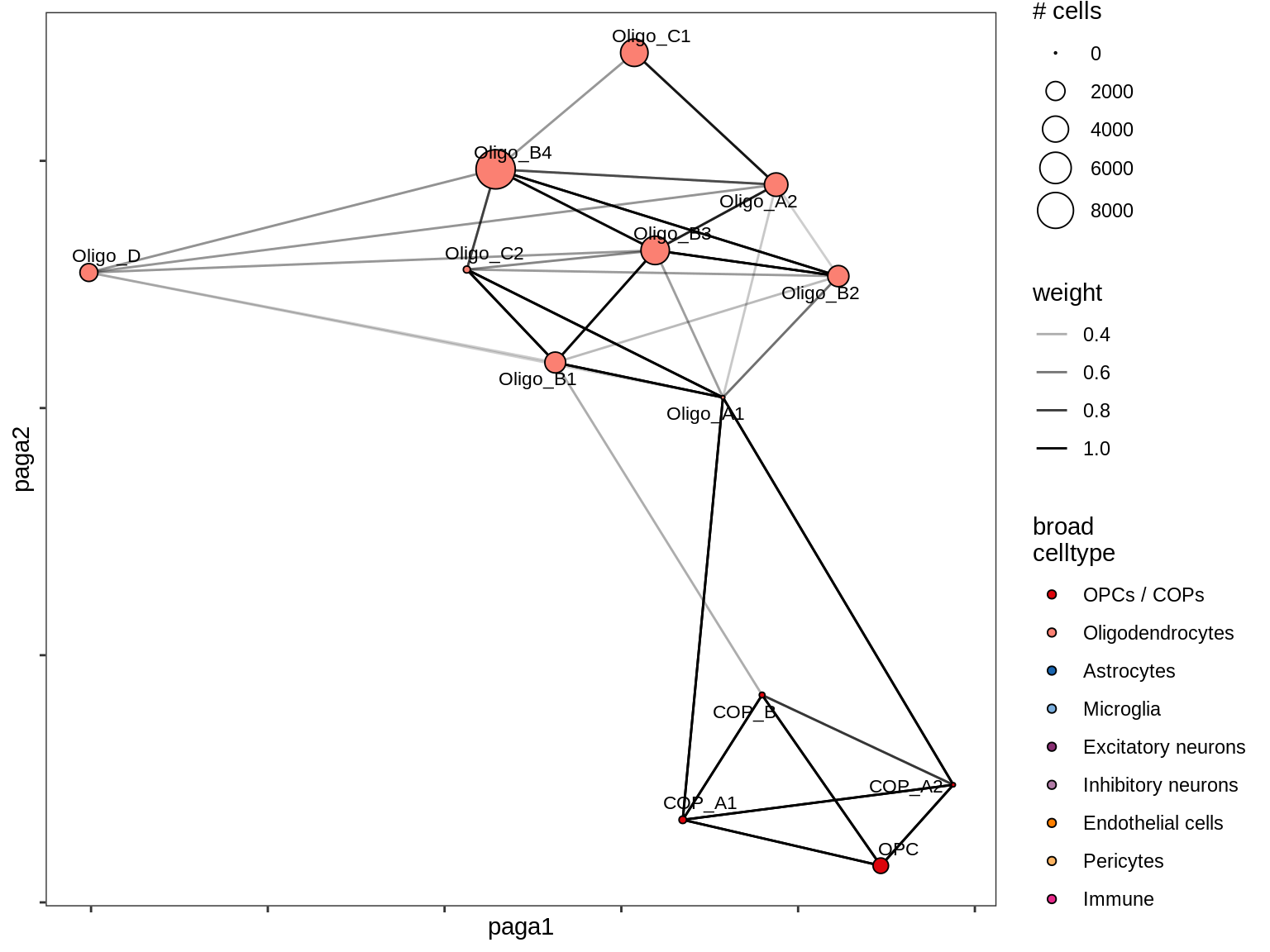
lesion_AL
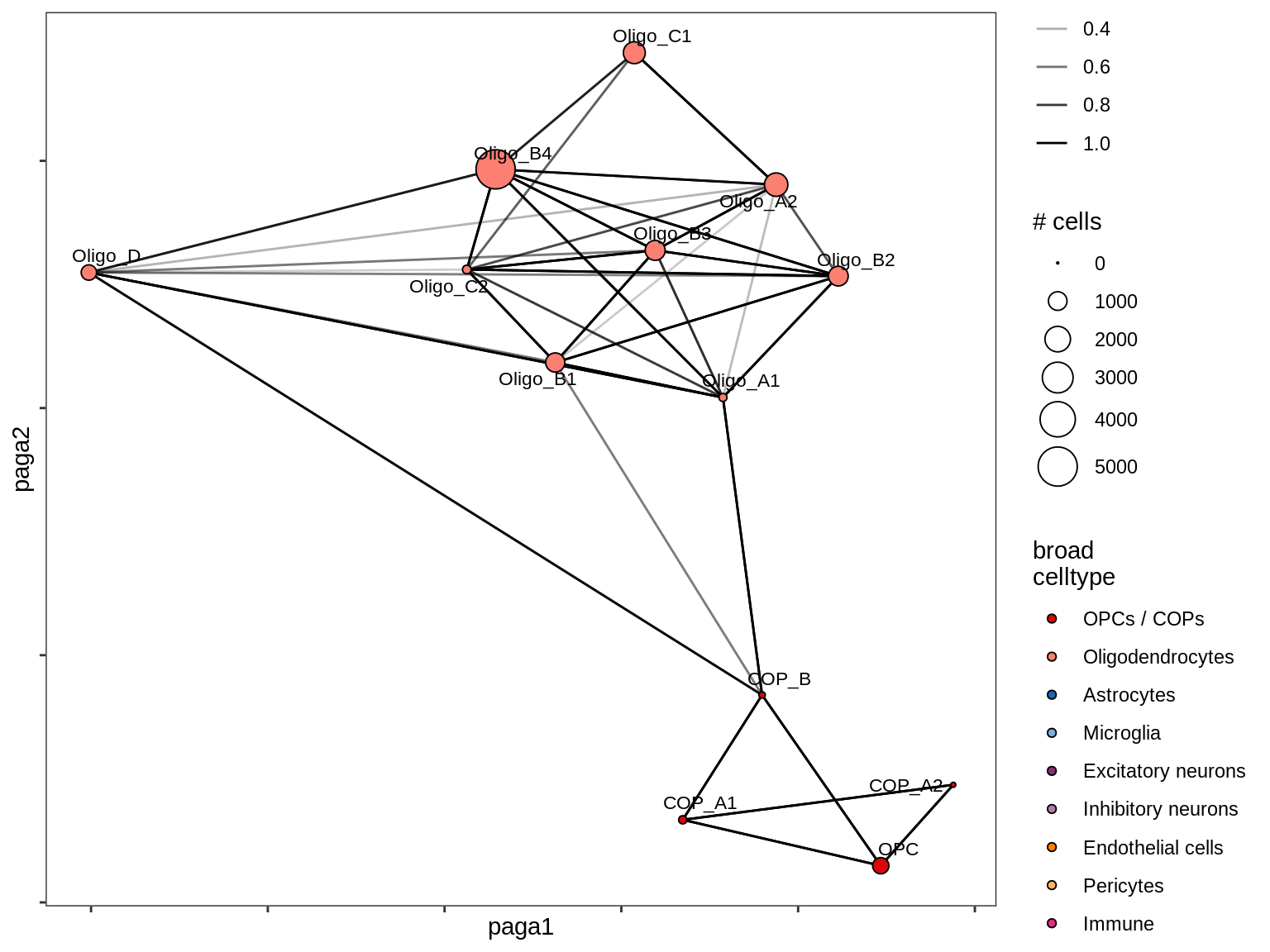
lesion_CAL
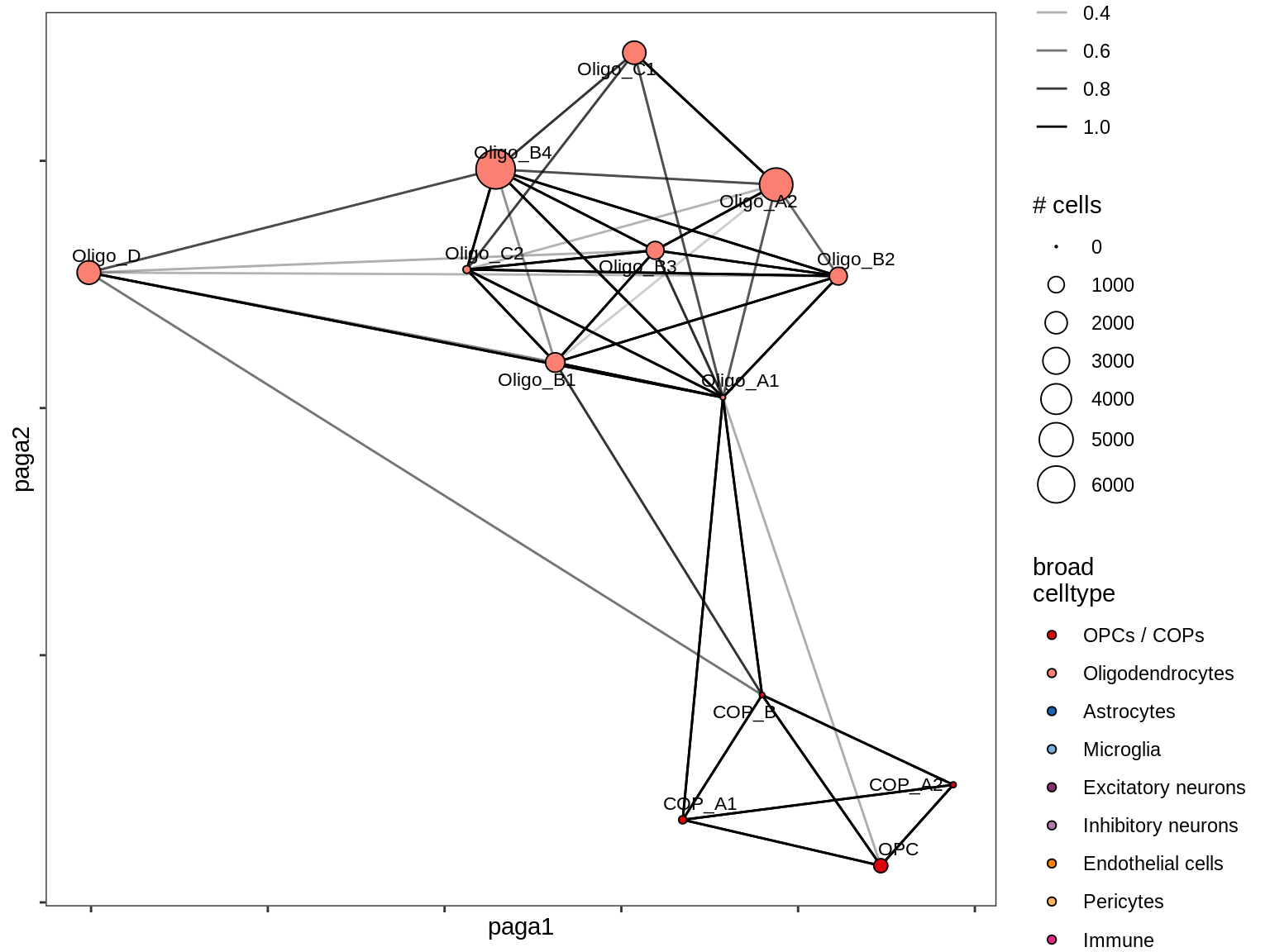
lesion_CIL
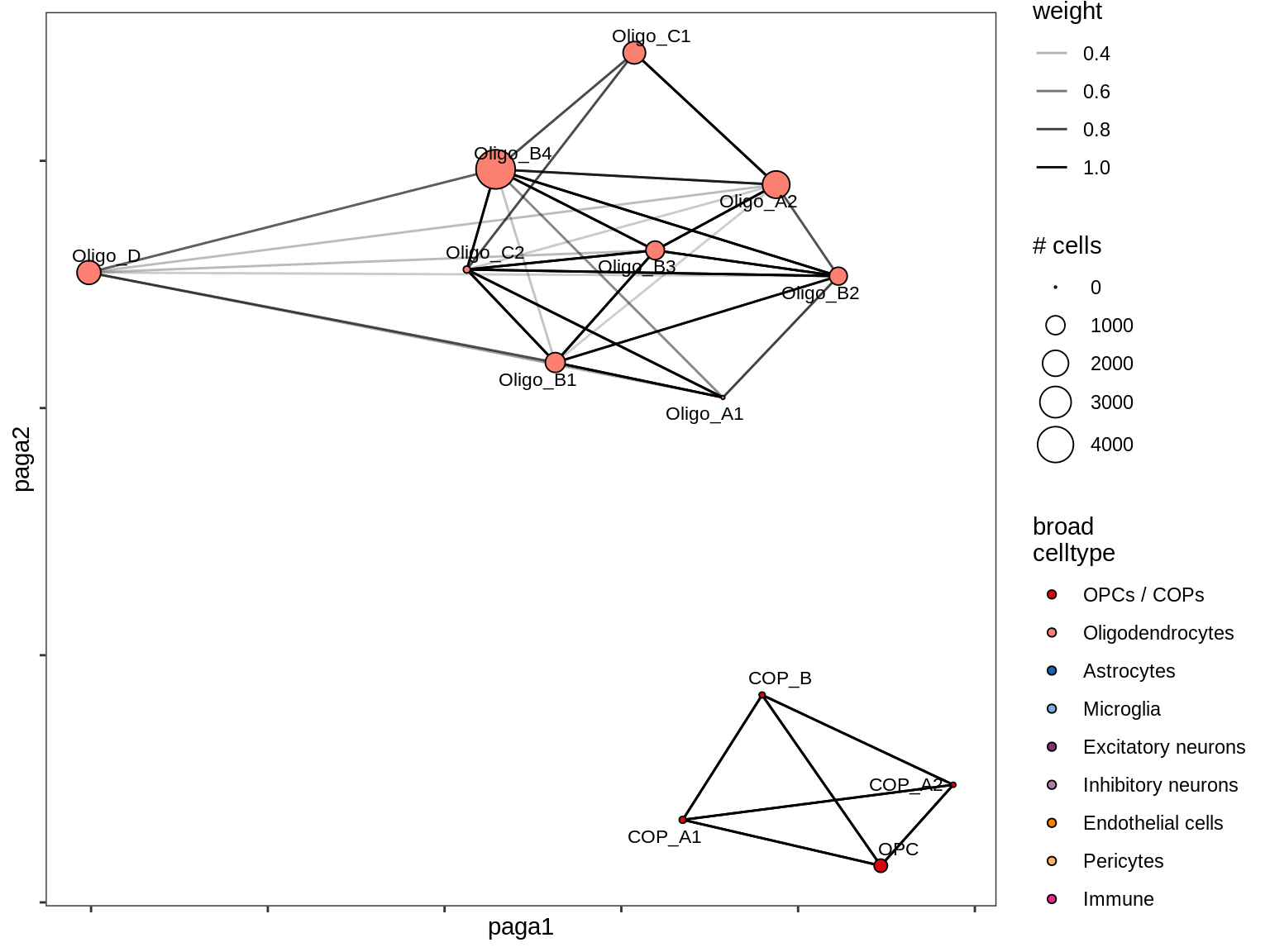
lesion_RL
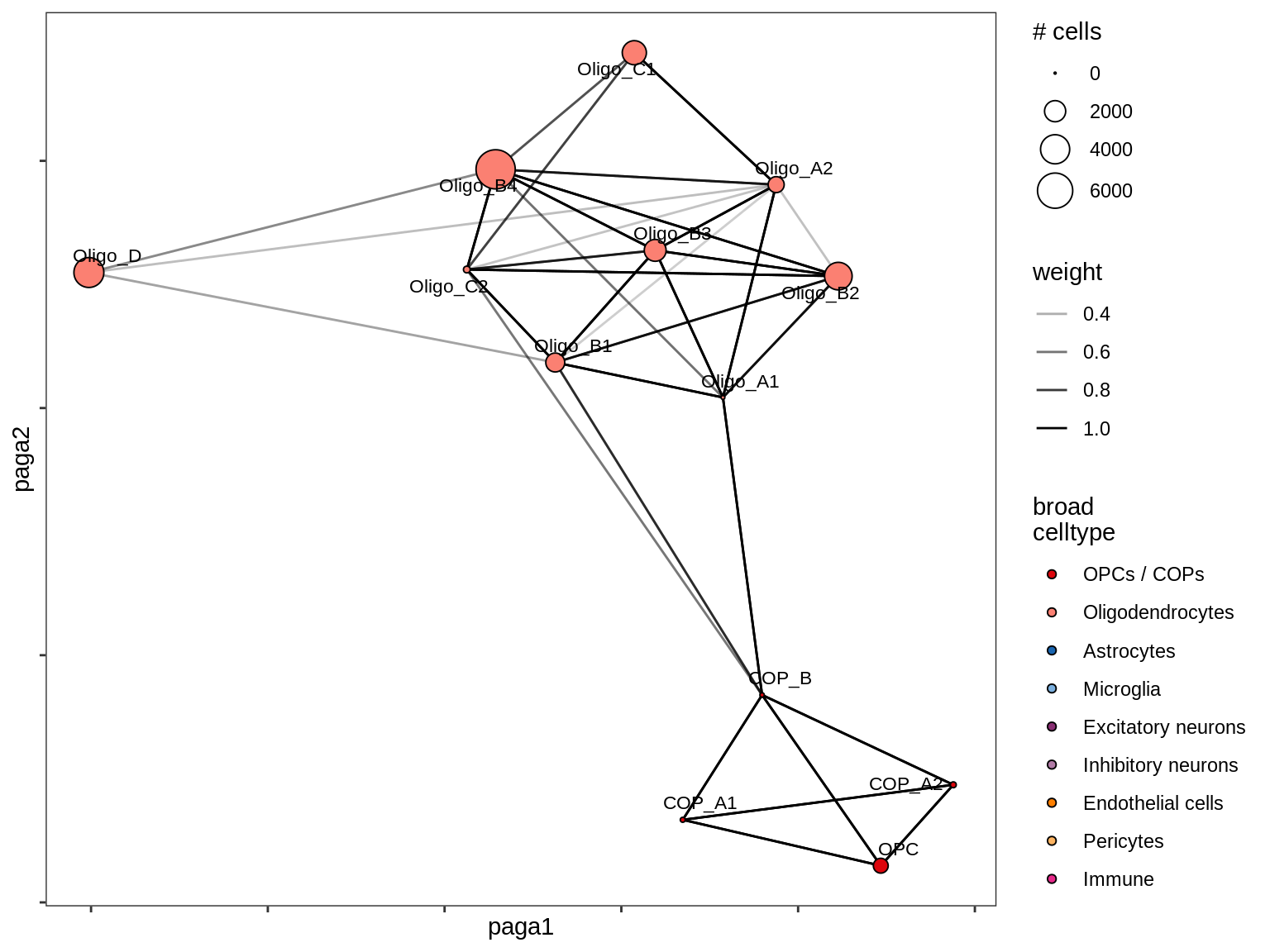
lesion_GM
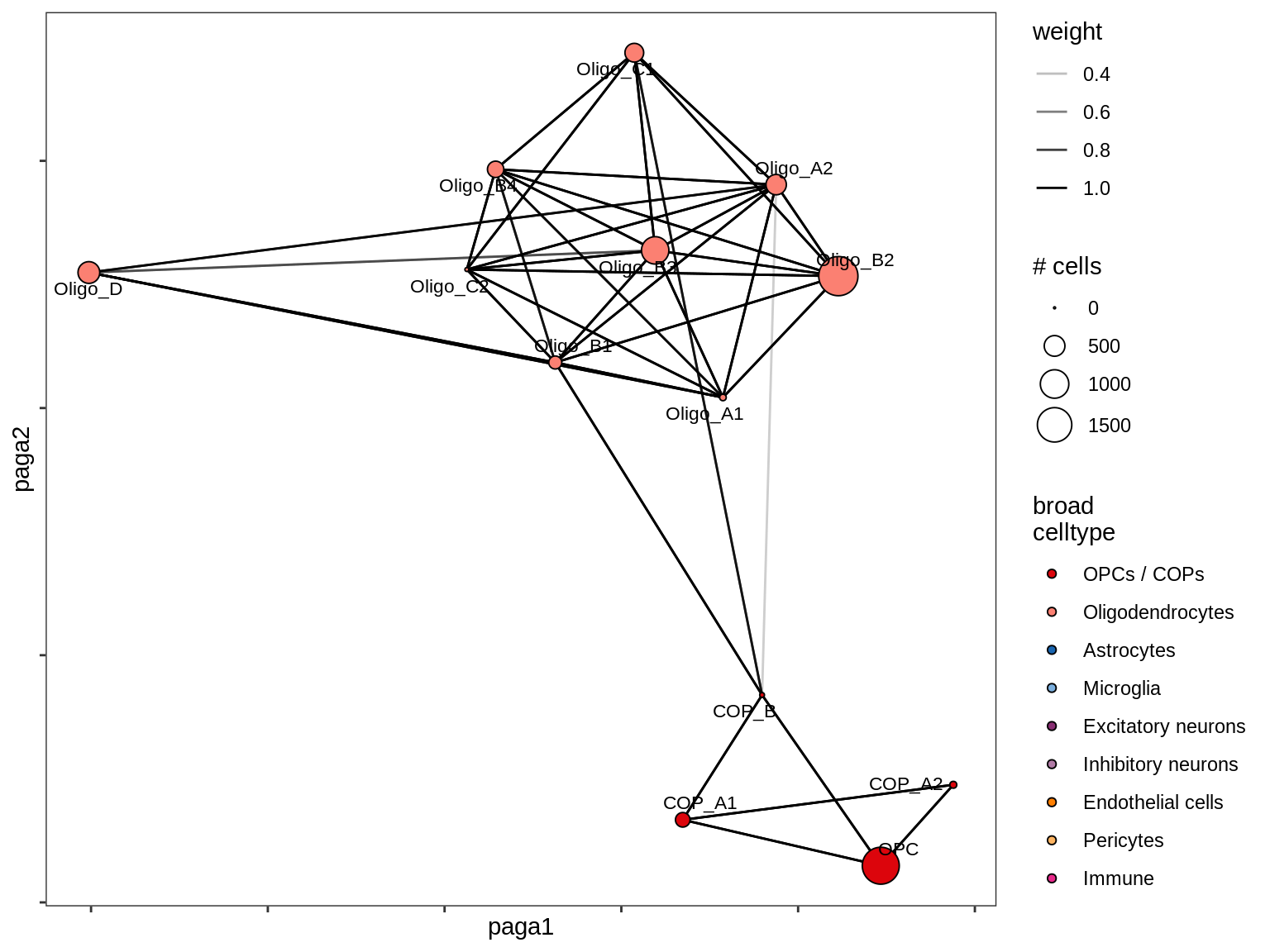
lesion_NAGM
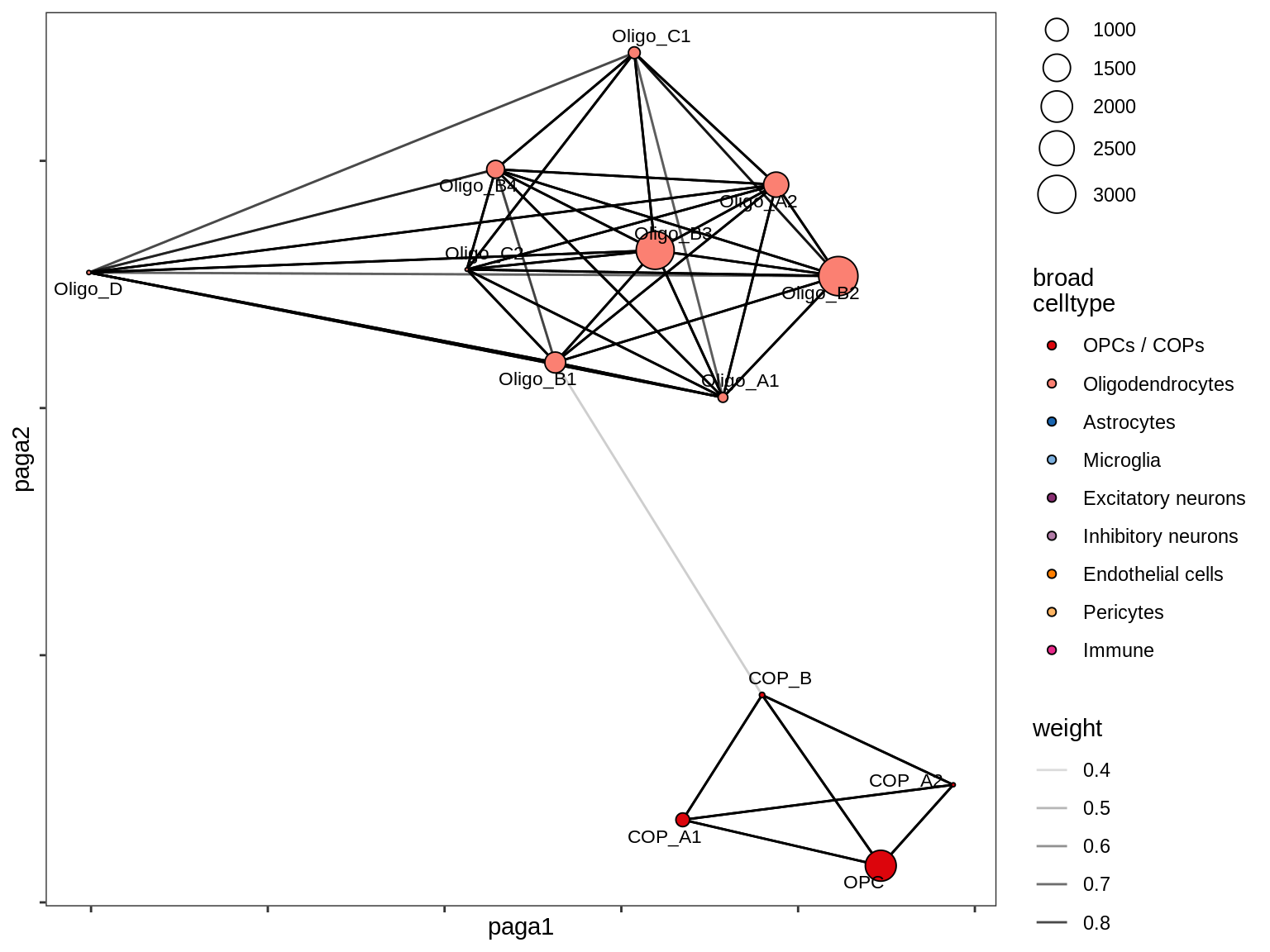
lesion_GML
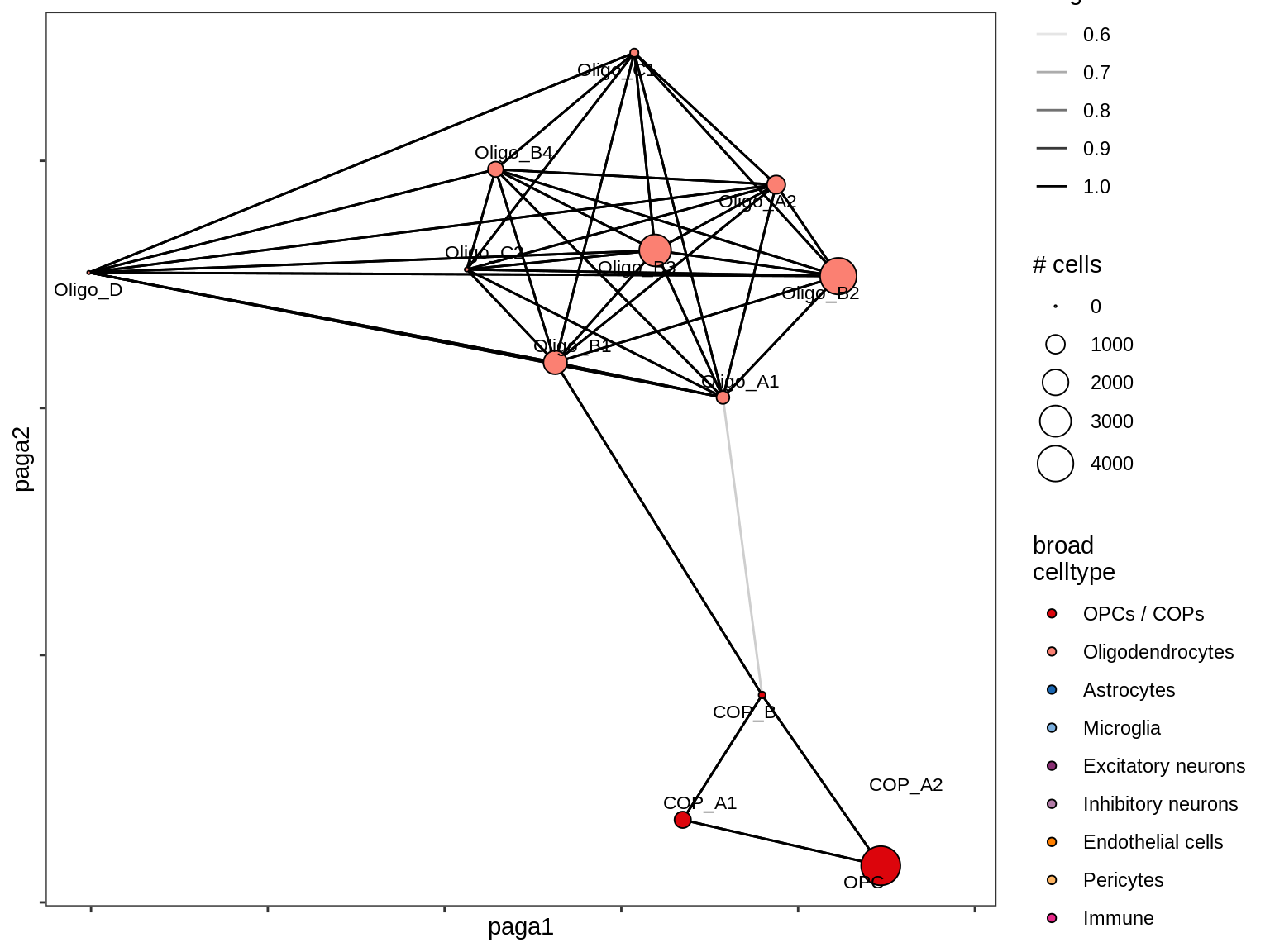
Split by lesion type, annotated by ANCOM coefficients
for (nn in names(lesion_list)) {
cat('#### ', nn, '\n')
ctrl_sample = xy_ref_list[[nn]] %>% str_match("(lesion|healthy)_(.+)") %>% .[3]
ancom_dt = ancom_ls[[ ctrl_sample ]]
suppressWarnings({print(plot_paga_w_ancom(save_dir, nn, date_tag, 'olg',
conos_dt, lesion_list[[nn]], labels_dt, xy_ref_n = "all",
ancom_dt, max_fc = 8))})
cat('\n\n')
}lesion_WM
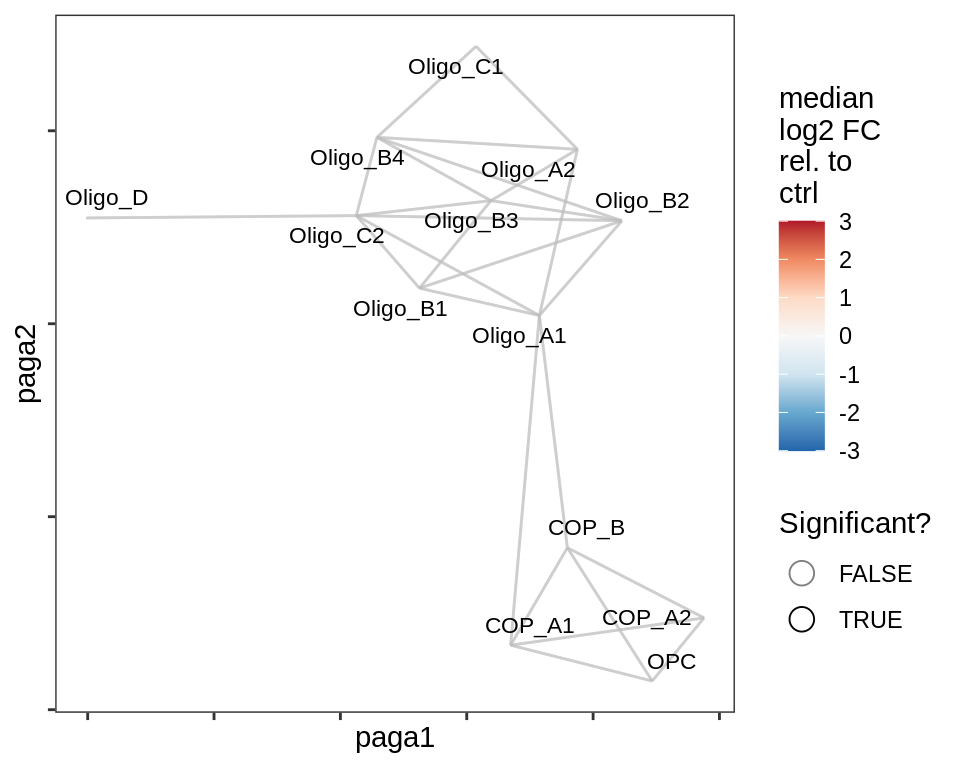
lesion_NAWM
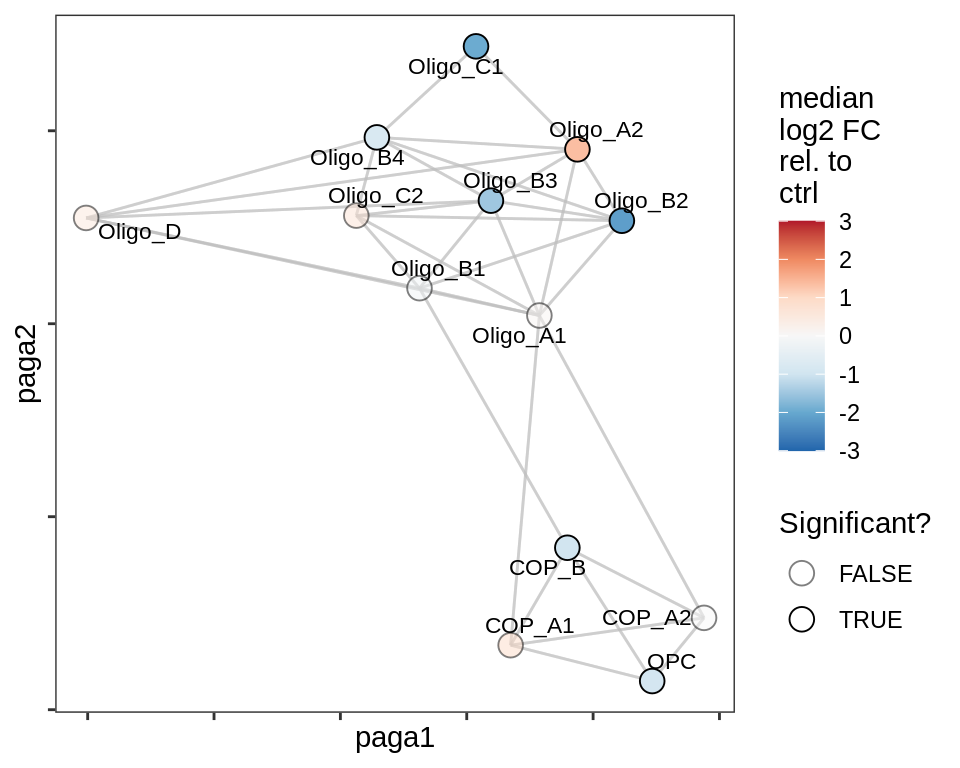
lesion_AL
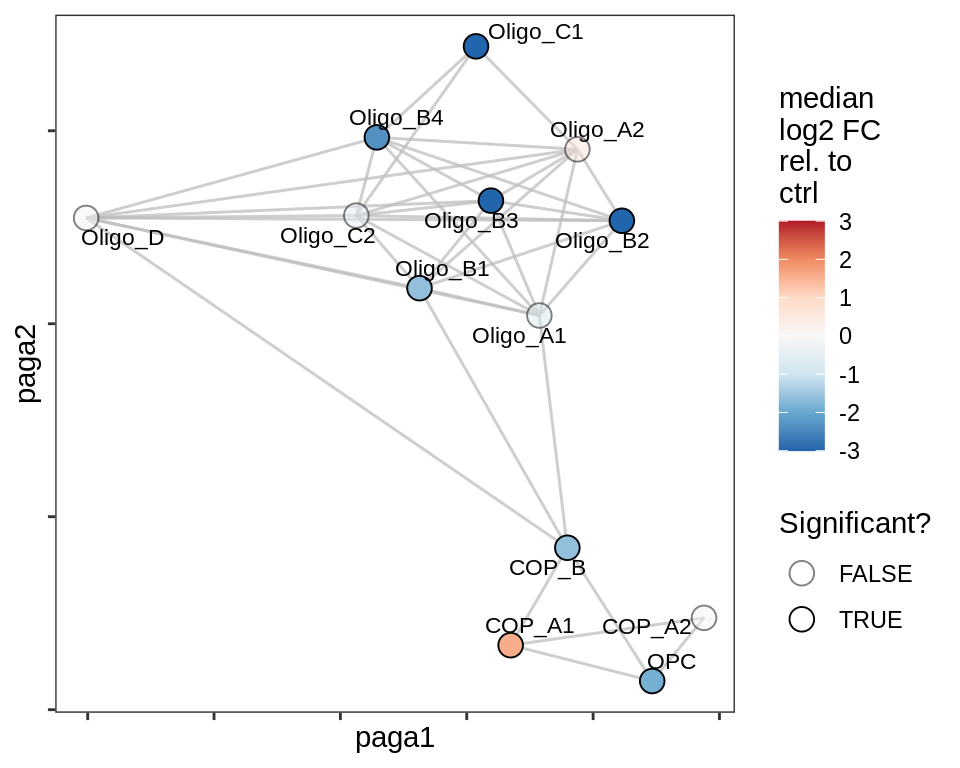
lesion_CAL
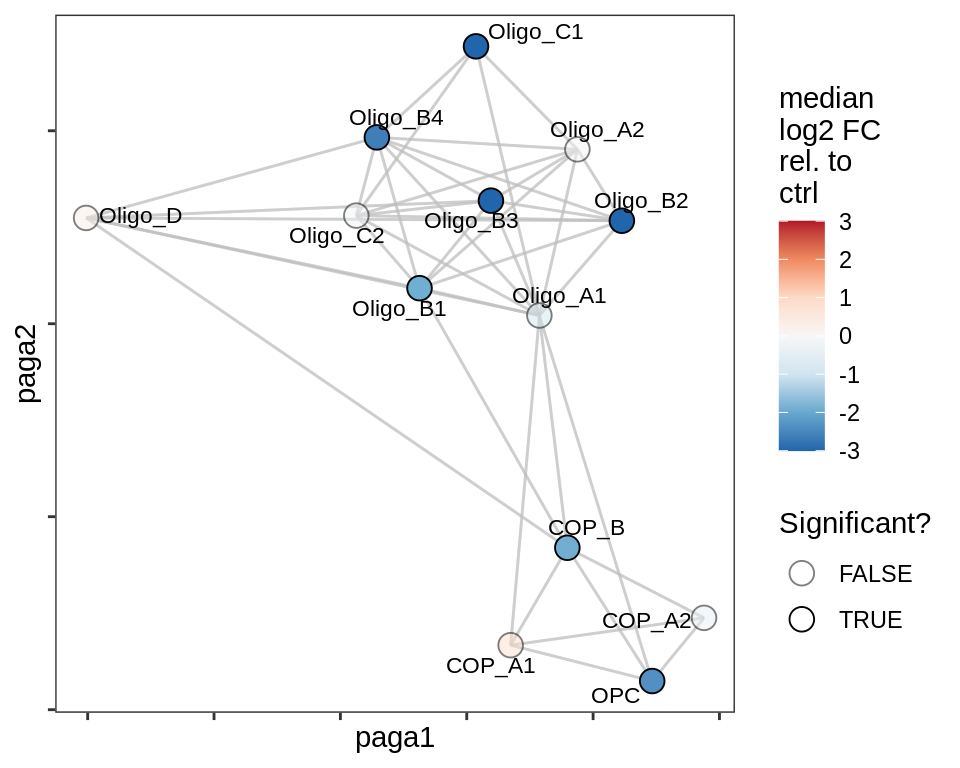
lesion_CIL
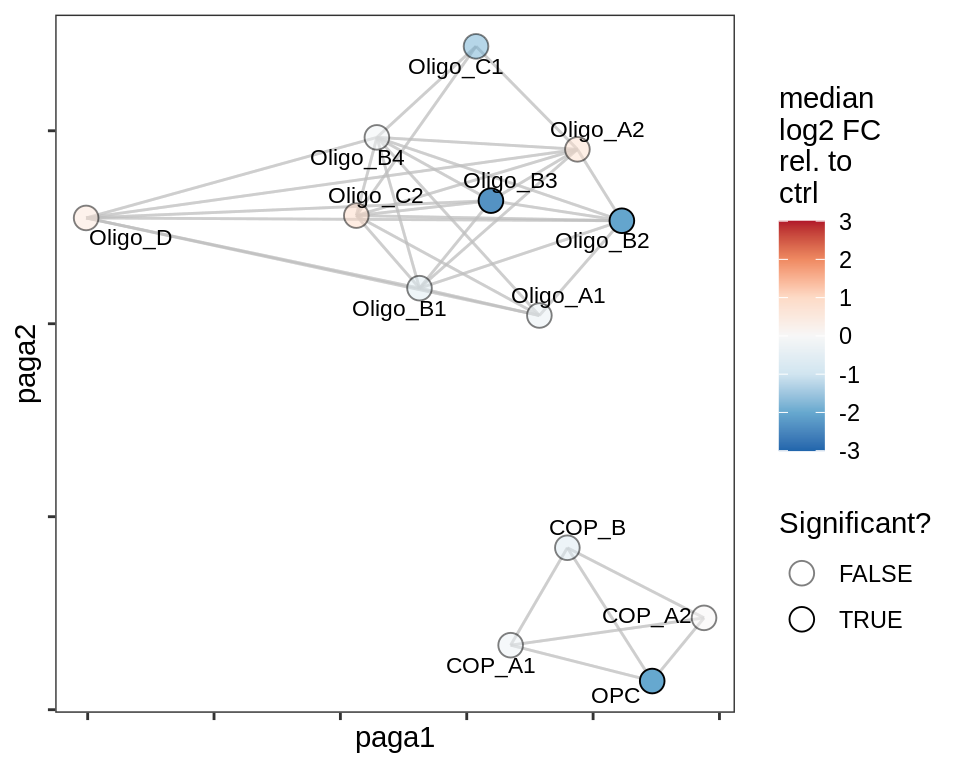
lesion_RL
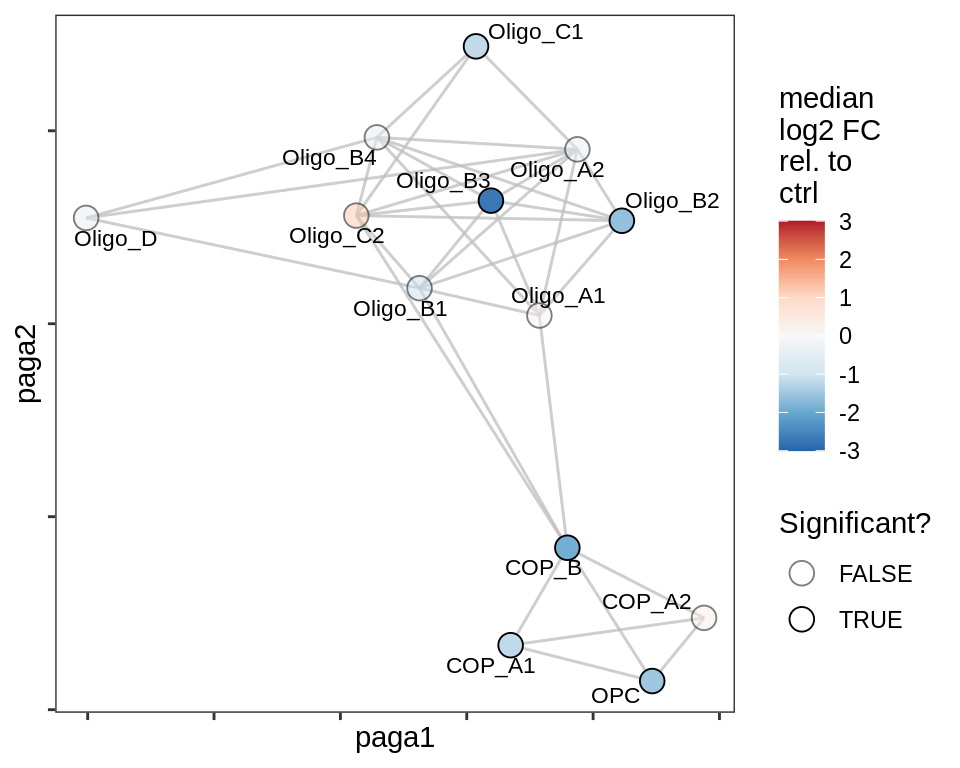
lesion_GM
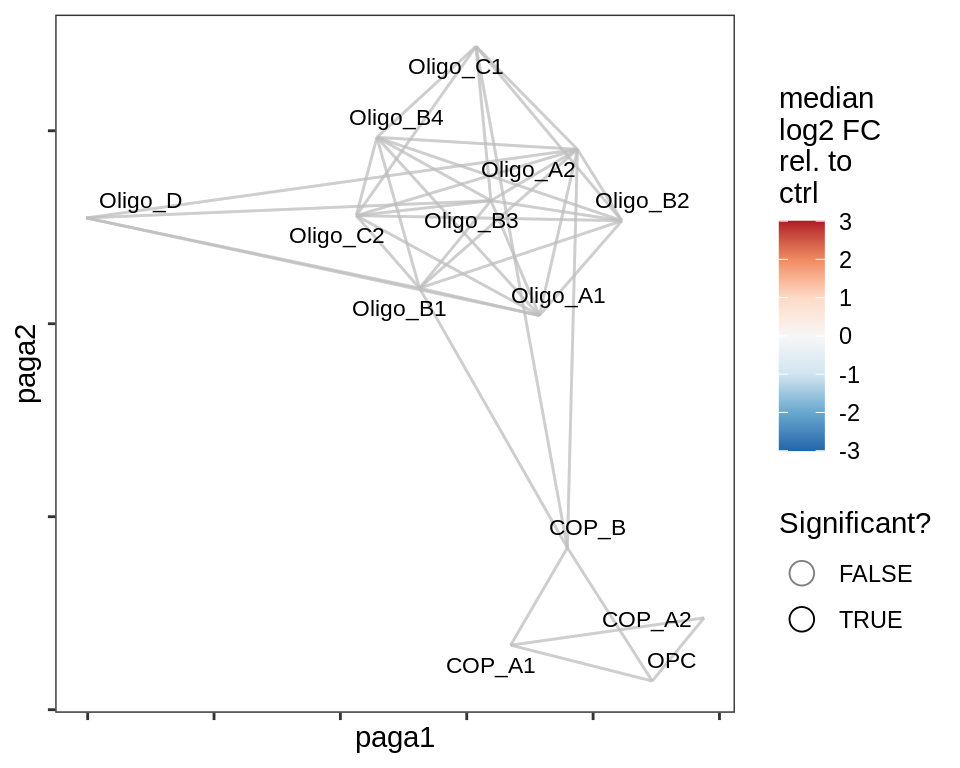
lesion_NAGM
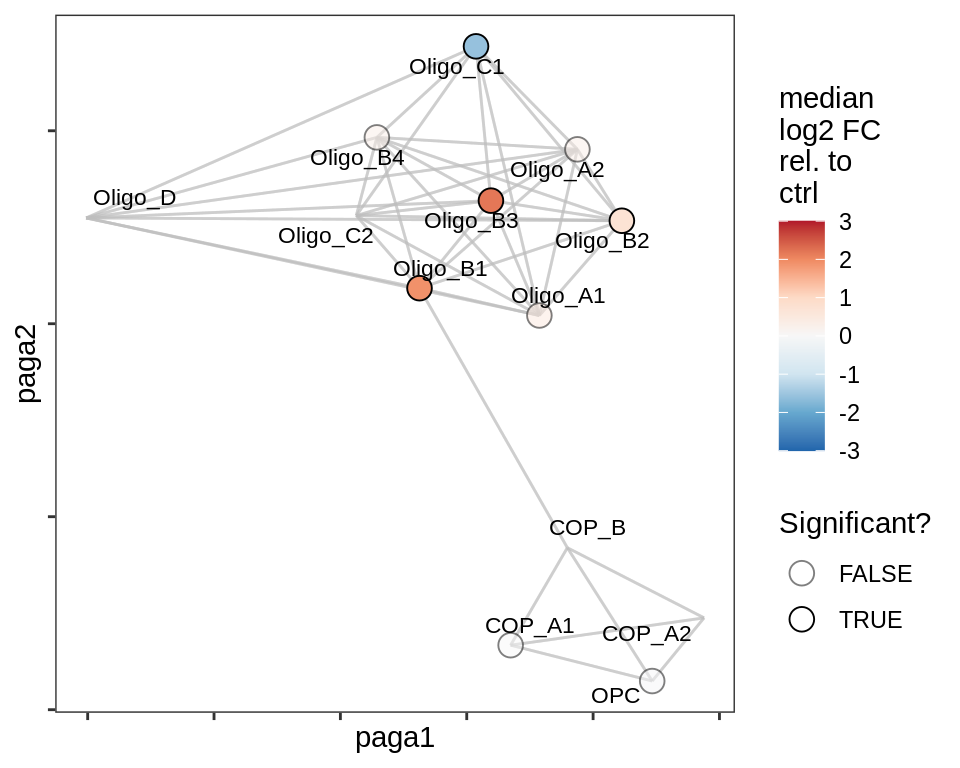
lesion_GML
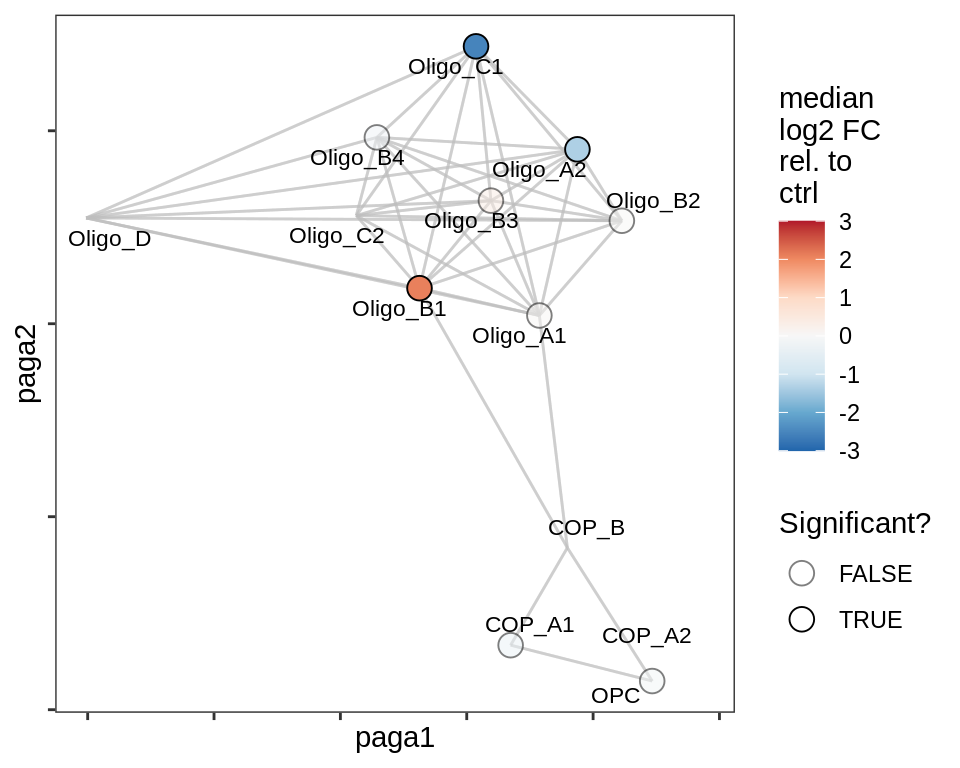
Heatmap of PAGA edges across samples, within oligo + OPC compartment
cat('### Data-driven')
draw(plot_edge_heatmap(edges_all, conos_dt, split = 'data'))
cat('\n')
cat('### Data-driven, no outliers')
draw(plot_edge_heatmap(edges_all, conos_dt[neuro_ok == TRUE], split = 'data'))
cat('\n')
cat('### Data-driven, WM no outliers')
draw(plot_edge_heatmap(edges_all, conos_dt[neuro_ok == TRUE & matter == 'WM'],
split = 'data'))
cat('\n')
cat('### Data-driven, GM no outliers')
draw(plot_edge_heatmap(edges_all, conos_dt[neuro_ok == TRUE & matter == 'GM'],
split = 'data'))
cat('\n')
cat('### Split by lesion')
draw(plot_edge_heatmap(edges_all, conos_dt, split = 'lesion_type'))
cat('\n')
cat('### Split by lesion, no outliers')
draw(plot_edge_heatmap(edges_all, conos_dt[neuro_ok == TRUE], split = 'lesion_type'))
cat('\n')Outputs
devtools::session_info()- Session info ---------------------------------------------------------------
setting value
version R version 4.0.5 (2021-03-31)
os CentOS Linux 7 (Core)
system x86_64, linux-gnu
ui X11
language (EN)
collate en_US.UTF-8
ctype C
tz Europe/Zurich
date 2022-01-05
- Packages -------------------------------------------------------------------
package * version date lib
ade4 1.7-18 2021-09-16 [1]
ANCOMBC * 1.0.5 2021-03-09 [1]
ape 5.5 2021-04-25 [1]
assertthat * 0.2.1 2019-03-21 [2]
beeswarm 0.4.0 2021-06-01 [1]
Biobase * 2.50.0 2020-10-27 [1]
BiocGenerics * 0.36.1 2021-04-16 [1]
BiocManager 1.30.16 2021-06-15 [1]
BiocParallel * 1.24.1 2020-11-06 [1]
BiocStyle * 2.18.1 2020-11-24 [1]
biomformat 1.18.0 2020-10-27 [1]
Biostrings 2.58.0 2020-10-27 [1]
bit 4.0.4 2020-08-04 [2]
bit64 4.0.5 2020-08-30 [2]
bitops 1.0-7 2021-04-24 [2]
bslib 0.3.1 2021-10-06 [2]
cachem 1.0.6 2021-08-19 [1]
Cairo 1.5-12.2 2020-07-07 [2]
callr 3.7.0 2021-04-20 [2]
cellranger 1.1.0 2016-07-27 [2]
circlize * 0.4.13 2021-06-09 [1]
cli 3.0.1 2021-07-17 [1]
clue 0.3-60 2021-10-11 [1]
cluster 2.1.2 2021-04-17 [2]
codetools 0.2-18 2020-11-04 [2]
colorout * 1.2-2 2021-04-15 [1]
colorspace 2.0-2 2021-06-24 [1]
ComplexHeatmap * 2.6.2 2020-11-12 [1]
crayon 1.4.1 2021-02-08 [2]
data.table * 1.14.2 2021-09-27 [2]
DBI 1.1.1 2021-01-15 [2]
DelayedArray 0.16.3 2021-03-24 [1]
desc 1.4.0 2021-09-28 [1]
devtools 2.4.2 2021-06-07 [1]
digest 0.6.28 2021-09-23 [2]
dplyr 1.0.7 2021-06-18 [2]
ellipsis 0.3.2 2021-04-29 [2]
evaluate 0.14 2019-05-28 [2]
fansi 0.5.0 2021-05-25 [2]
farver 2.1.0 2021-02-28 [2]
fastmap 1.1.0 2021-01-25 [2]
forcats * 0.5.1 2021-01-27 [2]
foreach 1.5.1 2020-10-15 [2]
fs 1.5.0 2020-07-31 [2]
generics 0.1.1 2021-10-25 [2]
GenomeInfoDb * 1.26.7 2021-04-08 [1]
GenomeInfoDbData 1.2.4 2021-04-15 [1]
GenomicRanges * 1.42.0 2020-10-27 [1]
GetoptLong 1.0.5 2020-12-15 [1]
ggbeeswarm * 0.6.0 2017-08-07 [1]
ggplot2 * 3.3.5 2021-06-25 [1]
ggrepel * 0.9.1 2021-01-15 [2]
git2r 0.28.0 2021-01-10 [1]
GlobalOptions 0.1.2 2020-06-10 [1]
glue 1.4.2 2020-08-27 [2]
gridExtra 2.3 2017-09-09 [2]
gtable 0.3.0 2019-03-25 [2]
gtools 3.9.2 2021-06-06 [2]
hdf5r * 1.3.3 2020-08-18 [2]
here 1.0.1 2020-12-13 [2]
highr 0.9 2021-04-16 [2]
htmltools 0.5.2 2021-08-25 [2]
httpuv 1.6.3 2021-09-09 [2]
ica * 1.0-2 2018-05-24 [2]
igraph 1.2.7 2021-10-15 [2]
IRanges * 2.24.1 2020-12-12 [1]
iterators * 1.0.13 2020-10-15 [2]
itertools * 0.1-3 2014-03-12 [1]
janitor 2.1.0 2021-01-05 [1]
jquerylib 0.1.4 2021-04-26 [2]
jsonlite 1.7.2 2020-12-09 [2]
kernlab 0.9-29 2019-11-12 [1]
knitr 1.36 2021-09-29 [1]
labeling 0.4.2 2020-10-20 [2]
later 1.3.0 2021-08-18 [2]
lattice 0.20-45 2021-09-22 [2]
lifecycle 1.0.1 2021-09-24 [2]
loomR * 0.2.0 2021-04-15 [1]
lubridate 1.8.0 2021-10-07 [2]
magrittr * 2.0.1 2020-11-17 [1]
MASS * 7.3-54 2021-05-03 [2]
Matrix * 1.3-4 2021-06-01 [2]
MatrixGenerics * 1.2.1 2021-01-30 [1]
matrixStats * 0.61.0 2021-09-17 [1]
mclust 5.4.7 2020-11-20 [1]
memoise 2.0.0 2021-01-26 [1]
mgcv 1.8-38 2021-10-06 [1]
microbiome 1.12.0 2020-10-27 [1]
mixtools 1.2.0 2020-02-07 [1]
multtest 2.46.0 2020-10-27 [1]
munsell 0.5.0 2018-06-12 [2]
mvnfast 0.2.7 2021-05-20 [1]
mvtnorm 1.1-3 2021-10-08 [1]
nlme 3.1-153 2021-09-07 [2]
nloptr 1.2.2.2 2020-07-02 [1]
patchwork * 1.1.1 2020-12-17 [2]
permute 0.9-5 2019-03-12 [1]
phyloseq * 1.34.0 2020-10-27 [1]
pillar 1.6.4 2021-10-18 [1]
pkgbuild 1.2.0 2020-12-15 [1]
pkgconfig 2.0.3 2019-09-22 [2]
pkgload 1.2.3 2021-10-13 [2]
plyr 1.8.6 2020-03-03 [2]
png 0.1-7 2013-12-03 [2]
prettyunits 1.1.1 2020-01-24 [2]
processx 3.5.2 2021-04-30 [2]
promises 1.2.0.1 2021-02-11 [2]
ps 1.6.0 2021-02-28 [2]
purrr * 0.3.4 2020-04-17 [2]
R.methodsS3 1.8.1 2020-08-26 [1]
R.oo 1.24.0 2020-08-26 [1]
R.utils 2.11.0 2021-09-26 [1]
R6 * 2.5.1 2021-08-19 [2]
rappdirs 0.3.3 2021-01-31 [2]
rbibutils 2.2.4 2021-10-11 [1]
RColorBrewer * 1.1-2 2014-12-07 [2]
Rcpp 1.0.7 2021-07-07 [1]
RCurl 1.98-1.5 2021-09-17 [1]
Rdpack 2.1.2 2021-06-01 [1]
readxl * 1.3.1 2019-03-13 [2]
remotes 2.4.1 2021-09-29 [1]
reshape2 1.4.4 2020-04-09 [2]
reticulate * 1.22 2021-09-17 [2]
rhdf5 2.34.0 2020-10-27 [1]
rhdf5filters 1.2.1 2021-05-03 [1]
Rhdf5lib 1.12.1 2021-01-26 [1]
rjson 0.2.20 2018-06-08 [1]
rlang 0.4.12 2021-10-18 [2]
rmarkdown 2.11 2021-09-14 [1]
rprojroot 2.0.2 2020-11-15 [2]
Rtsne 0.15 2018-11-10 [2]
S4Vectors * 0.28.1 2020-12-09 [1]
SampleQC * 0.6.1 2021-08-16 [1]
sass 0.4.0 2021-05-12 [2]
scales * 1.1.1 2020-05-11 [2]
segmented 1.3-4 2021-04-22 [1]
sessioninfo 1.1.1 2018-11-05 [1]
shape 1.4.6 2021-05-19 [1]
SingleCellExperiment * 1.12.0 2020-10-27 [1]
snakecase 0.11.0 2019-05-25 [1]
stringi 1.7.4 2021-08-25 [1]
stringr * 1.4.0 2019-02-10 [2]
SummarizedExperiment * 1.20.0 2020-10-27 [1]
survival 3.2-13 2021-08-24 [2]
testthat 3.1.0 2021-10-04 [2]
tibble 3.1.5 2021-09-30 [1]
tidyr 1.1.4 2021-09-27 [2]
tidyselect 1.1.1 2021-04-30 [2]
usethis 2.1.2 2021-10-25 [1]
utf8 1.2.2 2021-07-24 [1]
uwot 0.1.10 2020-12-15 [2]
vctrs 0.3.8 2021-04-29 [2]
vegan 2.5-7 2020-11-28 [1]
vipor 0.4.5 2017-03-22 [1]
viridis * 0.6.2 2021-10-13 [1]
viridisLite * 0.4.0 2021-04-13 [1]
withr 2.4.2 2021-04-18 [2]
workflowr * 1.6.2 2020-04-30 [1]
xfun 0.27 2021-10-18 [1]
XVector 0.30.0 2020-10-27 [1]
yaml 2.2.1 2020-02-01 [2]
zlibbioc 1.36.0 2020-10-27 [1]
source
CRAN (R 4.0.5)
Bioconductor
CRAN (R 4.0.3)
CRAN (R 4.0.0)
CRAN (R 4.0.3)
Bioconductor
Bioconductor
CRAN (R 4.0.3)
Bioconductor
Bioconductor
Bioconductor
Bioconductor
CRAN (R 4.0.2)
CRAN (R 4.0.2)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.5)
CRAN (R 4.0.2)
CRAN (R 4.0.3)
CRAN (R 4.0.0)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
Github (jalvesaq/colorout@79931fd)
CRAN (R 4.0.3)
Bioconductor
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
Bioconductor
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.0)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.2)
CRAN (R 4.0.5)
Bioconductor
Bioconductor
Bioconductor
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.0)
CRAN (R 4.0.0)
CRAN (R 4.0.3)
CRAN (R 4.0.2)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.5)
CRAN (R 4.0.0)
CRAN (R 4.0.5)
Bioconductor
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.5)
CRAN (R 4.0.5)
Github (mojaveazure/loomR@df0144b)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
Bioconductor
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
Bioconductor
CRAN (R 4.0.3)
Bioconductor
CRAN (R 4.0.0)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
Bioconductor
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.0)
CRAN (R 4.0.5)
CRAN (R 4.0.0)
CRAN (R 4.0.0)
CRAN (R 4.0.0)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.0)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.0)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.0)
CRAN (R 4.0.5)
CRAN (R 4.0.0)
CRAN (R 4.0.5)
Bioconductor
Bioconductor
Bioconductor
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.0)
Bioconductor
Github (wmacnair/SampleQC@3ae8a08)
CRAN (R 4.0.3)
CRAN (R 4.0.0)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.1)
Bioconductor
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.0)
Bioconductor
CRAN (R 4.0.5)
CRAN (R 4.0.5)
CRAN (R 4.0.5)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
CRAN (R 4.0.1)
CRAN (R 4.0.3)
CRAN (R 4.0.3)
CRAN (R 4.0.5)
Bioconductor
CRAN (R 4.0.3)
Bioconductor
[1] /pstore/home/macnairw/lib/conda_r3.12
[2] /pstore/home/macnairw/.conda/envs/r_4.0.3/lib/R/library
sessionInfo()R version 4.0.5 (2021-03-31)
Platform: x86_64-conda-linux-gnu (64-bit)
Running under: CentOS Linux 7 (Core)
Matrix products: default
BLAS/LAPACK: /pstore/home/macnairw/.conda/envs/r_4.0.3/lib/libopenblasp-r0.3.12.so
locale:
[1] LC_CTYPE=C LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] parallel stats4 grid stats graphics grDevices utils
[8] datasets methods base
other attached packages:
[1] SampleQC_0.6.1 SingleCellExperiment_1.12.0
[3] SummarizedExperiment_1.20.0 Biobase_2.50.0
[5] GenomicRanges_1.42.0 GenomeInfoDb_1.26.7
[7] IRanges_2.24.1 S4Vectors_0.28.1
[9] BiocGenerics_0.36.1 MatrixGenerics_1.2.1
[11] matrixStats_0.61.0 Matrix_1.3-4
[13] loomR_0.2.0 itertools_0.1-3
[15] iterators_1.0.13 hdf5r_1.3.3
[17] R6_2.5.1 BiocParallel_1.24.1
[19] ComplexHeatmap_2.6.2 ggbeeswarm_0.6.0
[21] ggrepel_0.9.1 reticulate_1.22
[23] MASS_7.3-54 phyloseq_1.34.0
[25] ANCOMBC_1.0.5 ica_1.0-2
[27] purrr_0.3.4 patchwork_1.1.1
[29] readxl_1.3.1 forcats_0.5.1
[31] ggplot2_3.3.5 scales_1.1.1
[33] viridis_0.6.2 viridisLite_0.4.0
[35] assertthat_0.2.1 stringr_1.4.0
[37] data.table_1.14.2 magrittr_2.0.1
[39] circlize_0.4.13 RColorBrewer_1.1-2
[41] BiocStyle_2.18.1 colorout_1.2-2
[43] workflowr_1.6.2
loaded via a namespace (and not attached):
[1] utf8_1.2.2 R.utils_2.11.0 tidyselect_1.1.1
[4] Rtsne_0.15 devtools_2.4.2 munsell_0.5.0
[7] codetools_0.2-18 withr_2.4.2 colorspace_2.0-2
[10] highr_0.9 knitr_1.36 Rdpack_2.1.2
[13] labeling_0.4.2 git2r_0.28.0 GenomeInfoDbData_1.2.4
[16] bit64_4.0.5 farver_2.1.0 rhdf5_2.34.0
[19] rprojroot_2.0.2 vctrs_0.3.8 generics_0.1.1
[22] xfun_0.27 clue_0.3-60 bitops_1.0-7
[25] rhdf5filters_1.2.1 microbiome_1.12.0 cachem_1.0.6
[28] DelayedArray_0.16.3 promises_1.2.0.1 beeswarm_0.4.0
[31] gtable_0.3.0 Cairo_1.5-12.2 processx_3.5.2
[34] rlang_0.4.12 GlobalOptions_0.1.2 splines_4.0.5
[37] BiocManager_1.30.16 yaml_2.2.1 reshape2_1.4.4
[40] httpuv_1.6.3 tools_4.0.5 usethis_2.1.2
[43] ellipsis_0.3.2 jquerylib_0.1.4 biomformat_1.18.0
[46] sessioninfo_1.1.1 Rcpp_1.0.7 plyr_1.8.6
[49] zlibbioc_1.36.0 RCurl_1.98-1.5 ps_1.6.0
[52] prettyunits_1.1.1 GetoptLong_1.0.5 cluster_2.1.2
[55] fs_1.5.0 here_1.0.1 mvnfast_0.2.7
[58] mvtnorm_1.1-3 pkgload_1.2.3 evaluate_0.14
[61] mclust_5.4.7 gridExtra_2.3 shape_1.4.6
[64] testthat_3.1.0 compiler_4.0.5 tibble_3.1.5
[67] crayon_1.4.1 R.oo_1.24.0 htmltools_0.5.2
[70] segmented_1.3-4 mgcv_1.8-38 later_1.3.0
[73] tidyr_1.1.4 lubridate_1.8.0 DBI_1.1.1
[76] rappdirs_0.3.3 ade4_1.7-18 permute_0.9-5
[79] cli_3.0.1 R.methodsS3_1.8.1 rbibutils_2.2.4
[82] igraph_1.2.7 pkgconfig_2.0.3 foreach_1.5.1
[85] vipor_0.4.5 bslib_0.3.1 multtest_2.46.0
[88] XVector_0.30.0 snakecase_0.11.0 callr_3.7.0
[91] digest_0.6.28 vegan_2.5-7 janitor_2.1.0
[94] Biostrings_2.58.0 rmarkdown_2.11 cellranger_1.1.0
[97] uwot_0.1.10 kernlab_0.9-29 gtools_3.9.2
[100] rjson_0.2.20 nloptr_1.2.2.2 lifecycle_1.0.1
[103] nlme_3.1-153 jsonlite_1.7.2 Rhdf5lib_1.12.1
[106] desc_1.4.0 fansi_0.5.0 pillar_1.6.4
[109] lattice_0.20-45 fastmap_1.1.0 pkgbuild_1.2.0
[112] survival_3.2-13 glue_1.4.2 remotes_2.4.1
[115] png_0.1-7 bit_4.0.4 stringi_1.7.4
[118] sass_0.4.0 mixtools_1.2.0 memoise_2.0.0
[121] dplyr_1.0.7 ape_5.5