Figures for manuscript
Will Macnair
Institute for Molecular Life Sciences, University of Zurich, SwitzerlandSwiss Institute of Bioinformatics (SIB), University of Zurich, SwitzerlandFebruary 02, 2022
Last updated: 2022-02-02
Checks: 4 3
Knit directory: MS_lesions/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of the R Markdown file created these results, you’ll want to first commit it to the Git repo. If you’re still working on the analysis, you can ignore this warning. When you’re finished, you can run wflow_publish to commit the R Markdown file and build the HTML.
The global environment had objects present when the code in the R Markdown file was run. These objects can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment. Use wflow_publish or wflow_build to ensure that the code is always run in an empty environment.
The following objects were defined in the global environment when these results were created:
| Name | Class | Size |
|---|---|---|
| q | function | 1008 bytes |
The command set.seed(20210118) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
- ancom_bootstrapped_gm_4_layer_pcs
- ancom_bootstrapped_gm_4pcs
- ancom_bootstrapped_gm
- ancom_bootstrapped_gm_no_layers
- ancom_bootstrapped_wm
- causes_of_variability_gm
- causes_of_variability_wm
- clr_plot_of_oligos
- cluster_mixing
- deg_barplot_gm
- deg_barplot_wm
- dotplot_marker_genes
- effect_of_including_pcs_neurons
- effect_of_including_pcs_others
- expression_heatmap_gm
- expression_heatmap_wm_clustered
- expression_heatmap_wm
- expression_heatmap_wm_lesions
- f1_vs_f2
- fig_mofa_factors_diagnosis
- fixed_vs_random_gm_4pcs
- fixed_vs_random_gm
- fixed_vs_random_wm
- gm_neuron_propns_layers
- gm_vs_wm_proportions
- grp17_cell_abundances
- gwas_coloc
- gwas_de_barplots
- gwas_manhattan
- module_scores_opc_oligo
- module_top_oligo_genes
- mofa_factor_heatmap_wm
- muscat_vs_sd
- oligo_barplot_gm
- oligo_barplot_wm
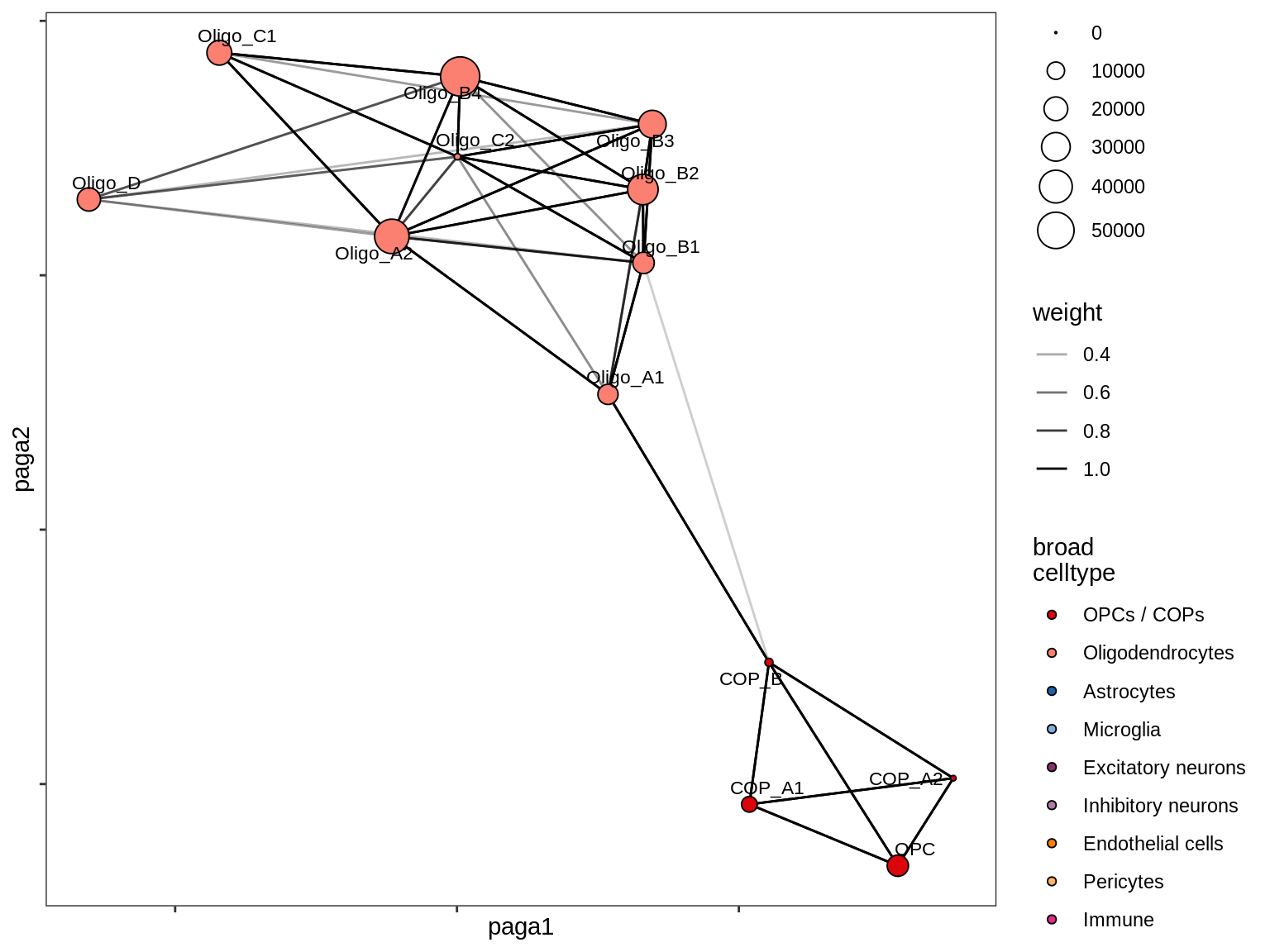
- paga_on_oligos
- random_effects_model_example
- sample_summary
- sccaf_summary
- session_info
- session-info-chunk-inserted-by-workflowr
- top_genes_factor1
- top_genes_factor2
- umap_all_celltypes
- umap_opc_oligo
- variance_explained
- wm_logfc_profile_clusters
To ensure reproducibility of the results, delete the cache directory ms99_manuscript_figures_cache and re-run the analysis. To have workflowr automatically delete the cache directory prior to building the file, set delete_cache = TRUE when running wflow_build() or wflow_publish().
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 965ed12. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rprofile
Ignored: .Rproj.user/
Ignored: ._MS_lesions.sublime-project
Ignored: ._broad_modules.png
Ignored: ._causes_of_variability.png
Ignored: ._check_module_gos.png
Ignored: ._fc_clusters.png
Ignored: ._logfc_hclust_dendro_astro.png
Ignored: ._olg_clr_kmeans.png
Ignored: ._paga_plot_test.png
Ignored: ._test_heatmap.png
Ignored: .log/
Ignored: MS_lesions.sublime-project
Ignored: MS_lesions.sublime-workspace
Ignored: analysis/.__site.yml
Ignored: analysis/fig_muscat_cache/
Ignored: analysis/ms00_manuscript_figures_cache/
Ignored: analysis/ms02_doublet_id_cache/
Ignored: analysis/ms03_SampleQC_cache/
Ignored: analysis/ms04_conos_cache/
Ignored: analysis/ms05_splitting_cache/
Ignored: analysis/ms06_sccaf_cache/
Ignored: analysis/ms07_soup_cache/
Ignored: analysis/ms08_modules_cache/
Ignored: analysis/ms08_modules_pseudobulk_cache/
Ignored: analysis/ms09_ancombc_cache/
Ignored: analysis/ms09_ancombc_clean_1e3_cache/
Ignored: analysis/ms09_ancombc_clean_2e3_cache/
Ignored: analysis/ms09_ancombc_mixed_cache/
Ignored: analysis/ms10_muscat_run01_cache/
Ignored: analysis/ms10_muscat_run02_cache/
Ignored: analysis/ms10_muscat_template_broad_slim_cache/
Ignored: analysis/ms10_muscat_template_fine_slim_cache/
Ignored: analysis/ms11_paga_cache/
Ignored: analysis/ms11_paga_recalc_cache/
Ignored: analysis/ms11_paga_superclean_cache/
Ignored: analysis/ms12_markers_cache/
Ignored: analysis/ms13_labelling_cache/
Ignored: analysis/ms14_lesions_cache/
Ignored: analysis/ms15_mofa_sample_gm_cache/
Ignored: analysis/ms15_mofa_sample_gm_final_meta_cache/
Ignored: analysis/ms15_mofa_sample_gm_superclean_cache/
Ignored: analysis/ms15_mofa_sample_gm_w_layers_final_meta_cache/
Ignored: analysis/ms15_mofa_sample_wm_cache/
Ignored: analysis/ms15_mofa_sample_wm_final_meta_bigger_cache/
Ignored: analysis/ms15_mofa_sample_wm_final_meta_cache/
Ignored: analysis/ms15_mofa_sample_wm_new_meta_cache/
Ignored: analysis/ms15_mofa_sample_wm_superclean_cache/
Ignored: analysis/ms15_patients_cache/
Ignored: analysis/ms15_patients_gm_cache/
Ignored: analysis/ms15_patients_sample_level_cache/
Ignored: analysis/ms15_patients_w_ms_cache/
Ignored: analysis/ms99_deg_figures_gm_cache/
Ignored: analysis/ms99_deg_figures_wm_cache/
Ignored: analysis/ms99_manuscript_figures_cache/
Ignored: analysis/supp06_sccaf_cache/
Ignored: analysis/supp07_superclean_check_cache/
Ignored: analysis/supp09_ancombc_cache/
Ignored: analysis/supp09_ancombc_mixed_cache/
Ignored: analysis/supp09_ancombc_rowitch_cache/
Ignored: analysis/supp09_ancombc_superclean_cache/
Ignored: analysis/supp10_muscat_cache/
Ignored: analysis/supp10_muscat_ctrl_gm_vs_wm_cache/
Ignored: analysis/supp10_muscat_gm_layers_effects_cache/
Ignored: analysis/supp10_muscat_gsea_cache/
Ignored: analysis/supp10_muscat_heatmaps_cache/
Ignored: analysis/supp10_muscat_olg_pc1_cache/
Ignored: analysis/supp10_muscat_olg_pc2_cache/
Ignored: analysis/supp10_muscat_olg_pc_cache/
Ignored: analysis/supp10_muscat_regression_cache/
Ignored: analysis/supp10_muscat_soup_cache/
Ignored: analysis/supp10_muscat_soup_mito_cache/
Ignored: code/._ms10_muscat_fns_recover.R
Ignored: code/.recovery/
Ignored: code/jobs/._muscat_run09_2021-10-11.slurm
Ignored: code/muscat_plan.txt
Ignored: data/
Ignored: figures/
Ignored: output/
Ignored: tmp/
Untracked files:
Untracked: analysis/ms16_wgcna.R
Untracked: code/dev_dheeraj_oligo_dotplots_2021-12-08.R
Untracked: code/dev_magma_de_heatmaps_2021-12-03.R
Untracked: code/dev_plot_dotplot_sel_genes_2021-12-21.R
Untracked: code/dev_plot_fcs_selected_genes_2021-12-09.R
Untracked: code/tmp_compare_fgsea_vs_metascape.R
Untracked: logfc_hclust_dendro_astro.png
Untracked: olg_clr_kmeans.png
Unstaged changes:
Modified: analysis/ms09_ancombc_mixed.Rmd
Modified: analysis/ms13_labelling.Rmd
Modified: analysis/ms14_lesions.Rmd
Modified: analysis/ms15_mofa_sample_gm_w_layers_final_meta.Rmd
Modified: analysis/ms15_mofa_sample_wm_final_meta.Rmd
Modified: analysis/ms15_mofa_sample_wm_final_meta_bigger.Rmd
Modified: analysis/ms99_deg_figures_gm.Rmd
Modified: analysis/ms99_deg_figures_wm.Rmd
Modified: analysis/ms99_manuscript_figures.Rmd
Modified: code/dev_de_w_contamation_2021-10-25.R
Modified: code/dev_edger_on_mofa_20210804.R
Modified: code/ms03_SampleQC.R
Modified: code/ms09_ancombc_mixed.R
Modified: code/ms10_muscat_fns.R
Modified: code/ms14_lesions.R
Modified: code/ms99_deg_figures.R
Modified: code/supp10_muscat.R
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were made to the R Markdown (analysis/ms99_manuscript_figures.Rmd) and HTML (public/ms99_manuscript_figures.html) files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | fea6733 | wmacnair | 2022-01-27 | Update location of manuscript figures |
| html | fea6733 | wmacnair | 2022-01-27 | Update location of manuscript figures |
Setup / definitions
Libraries
Helper functions
source('code/ms00_utils.R')
library("knitr")Figure 1
B
UMAP applied to subset of 100k cells (subset because of memory limits), using parameters min_dist = 1, spread = 2, otherwise defaults. Clusters are determined by Louvain clustering applied to the conos graph, followed by post-hoc splitting of two clusters based on biological expectations (COPs and immune cells), and merging of very similar clusters (using SCCAF).
include_graphics("figure/ms12_markers.Rmd/plot_umap_final_celltypes_sel-1.png", error = FALSE)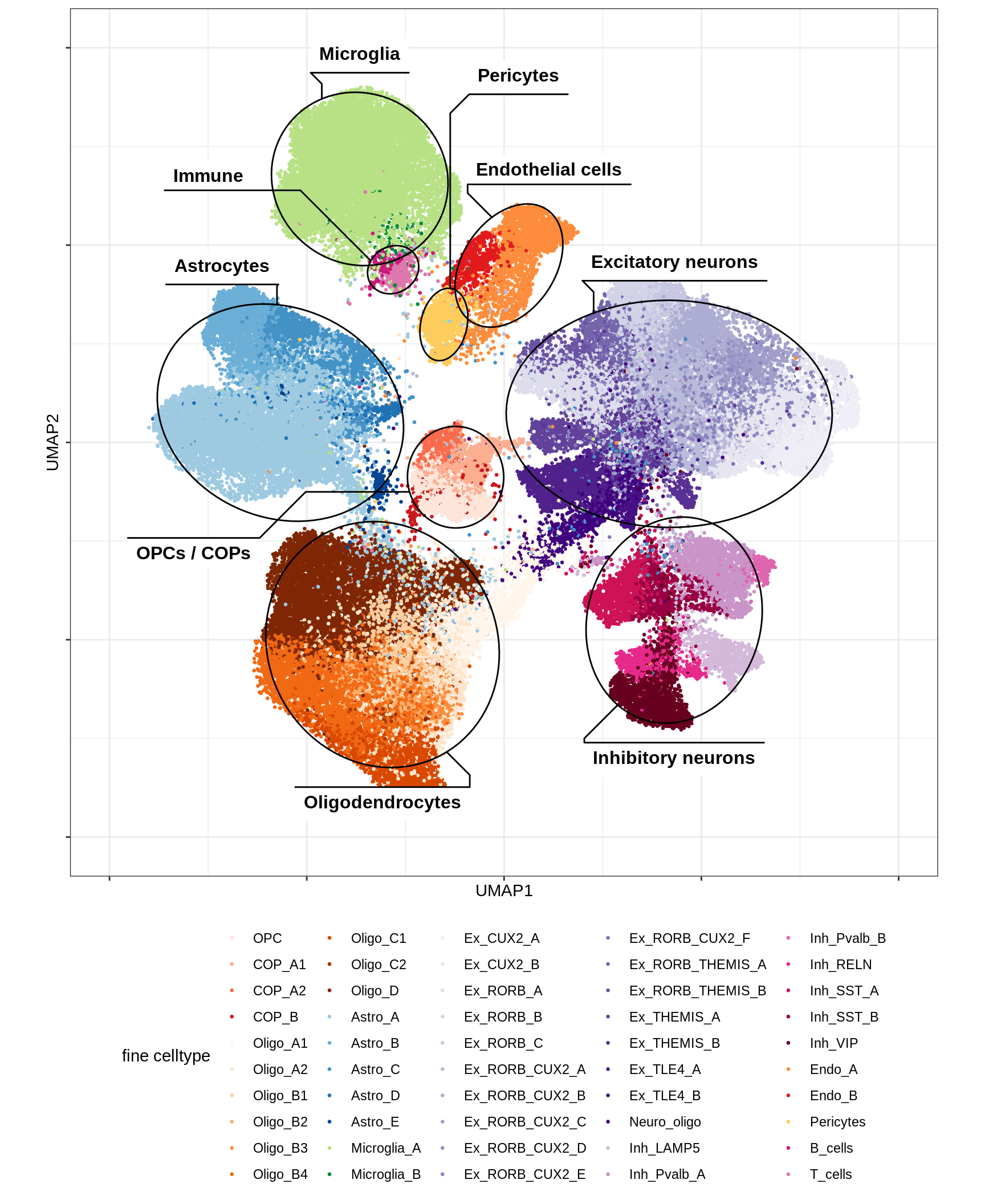
| Version | Author | Date |
|---|---|---|
| 7fb1b95 | wmacnair | 2021-11-25 |
C
UMAP plot as in B-1, restricted to just oligodendrocyte and OPC celltypes.
include_graphics("figure/ms12_markers.Rmd/plot_umap_opc_oligo_only-1.png", error = FALSE)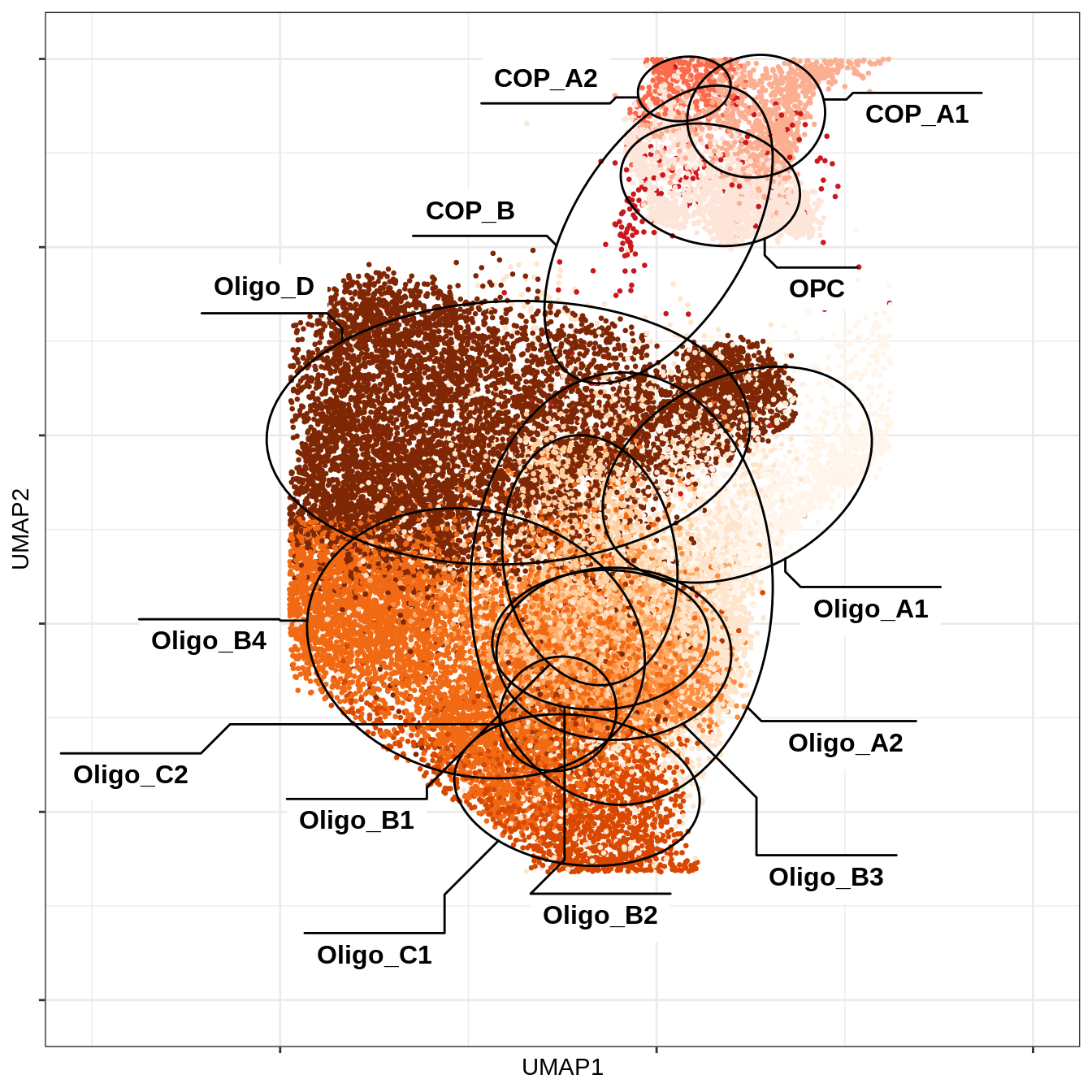
| Version | Author | Date |
|---|---|---|
| 7fb1b95 | wmacnair | 2021-11-25 |
D
Median oNMF module score per fine celltype for OPC and oligo modules and cells. Columns are scaled to have max value equal to 1.
include_graphics("figure/ms08_modules.Rmd/plot_scores_by_type_scaled-1.png", error = FALSE)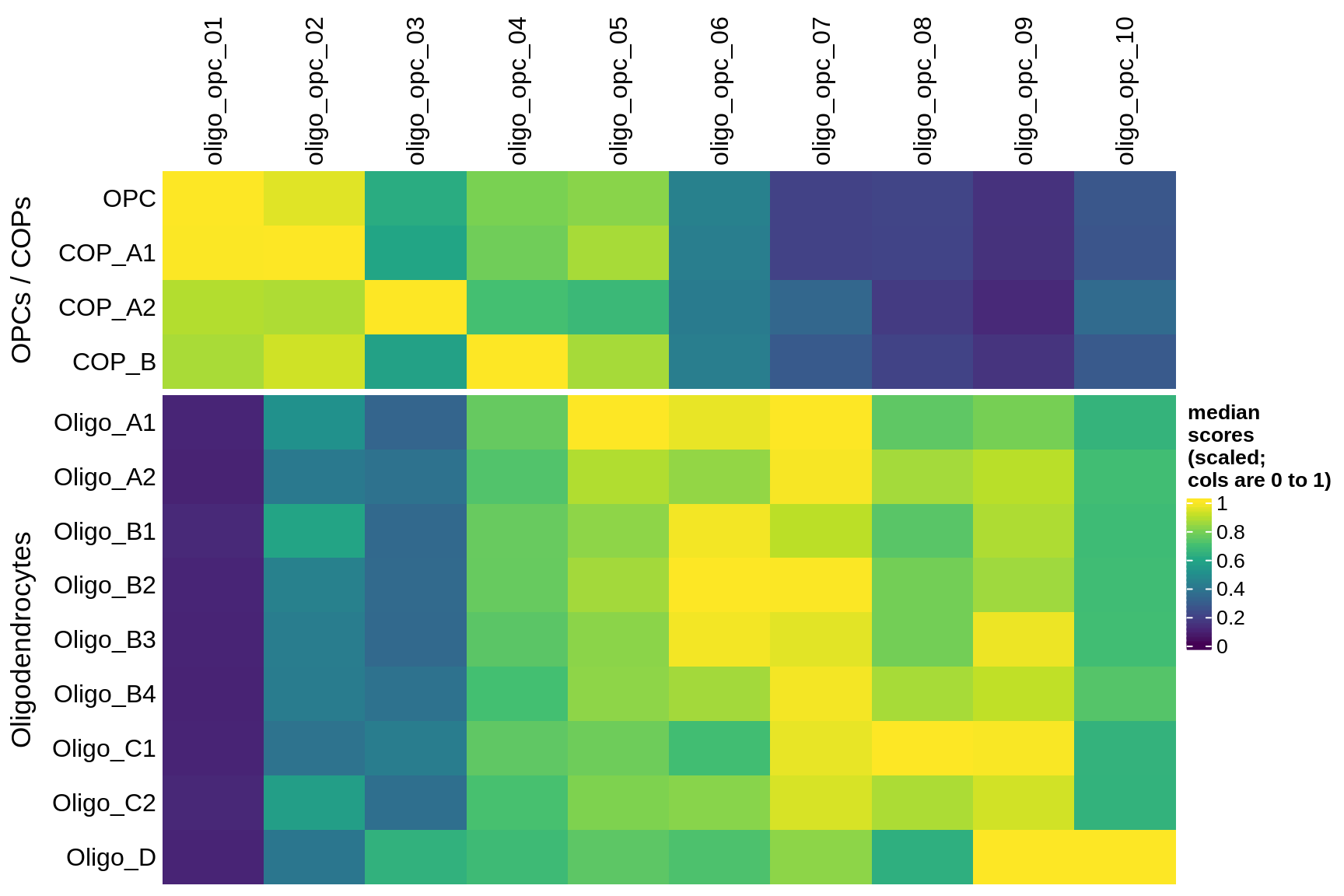
E
Expression of top genes for each oligo-OPC module (gene selected if weight >2%). Expression calculated across all cells and samples.
include_graphics("figure/ms08_modules.Rmd/plot_genes_dotplot-2.png", error = FALSE)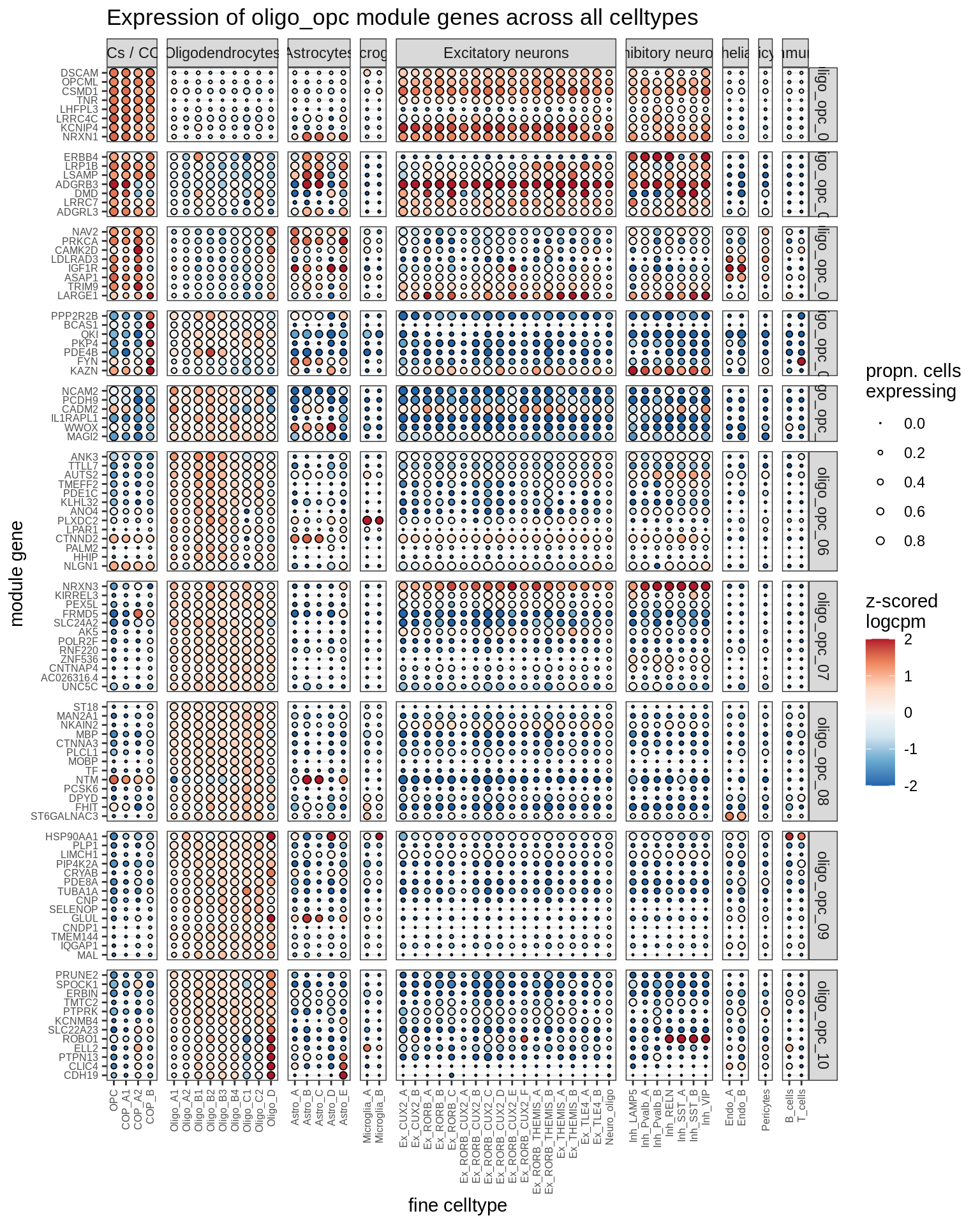
Figure 2
A
Proportions of fine celltypes in healthy GM and healthy WM. Neuronal celltypes excluded. Negative binomial model fit to absolute numbers for each celltype, using total number of cells in sample as offset. FDR calculated across all celltypes.
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_wm_vs_gm-1.png", error = FALSE)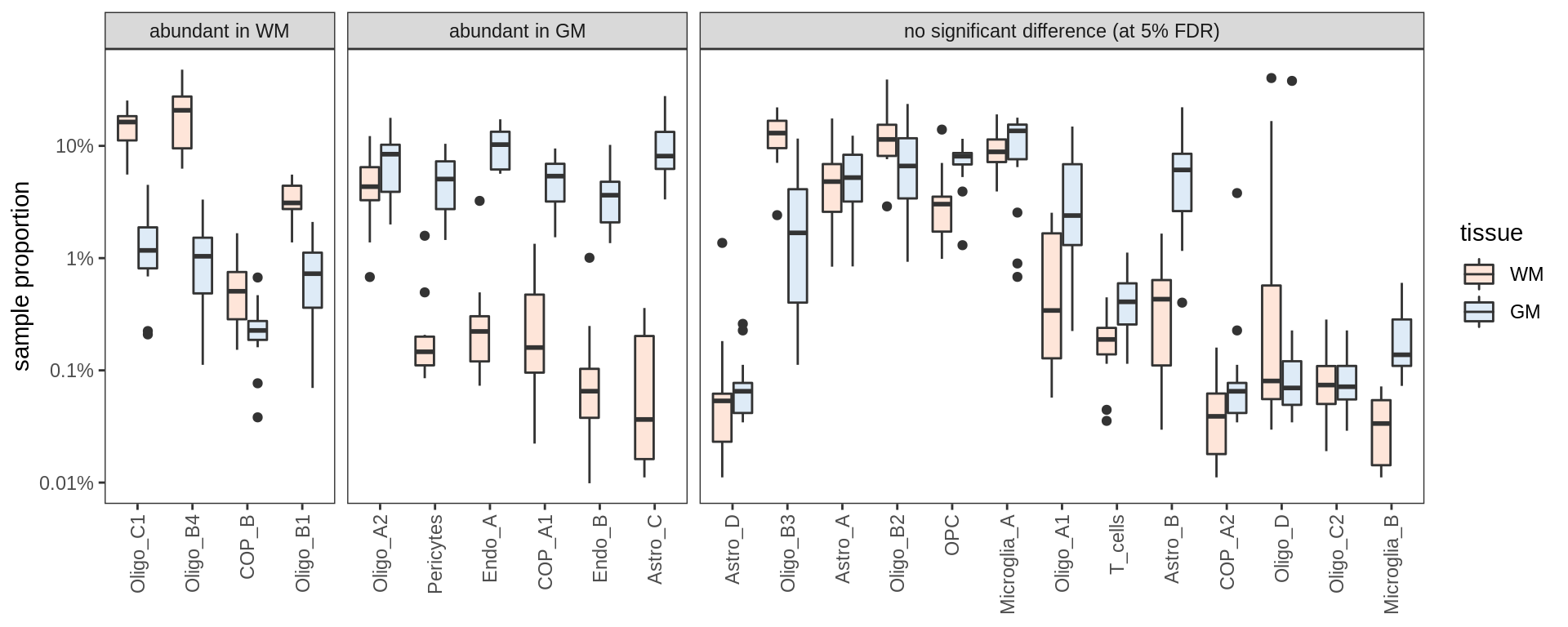
B
Contribution to variability in celltype abundances explained by lesion + patient in WM.
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_lrt_results-1.png", error = FALSE)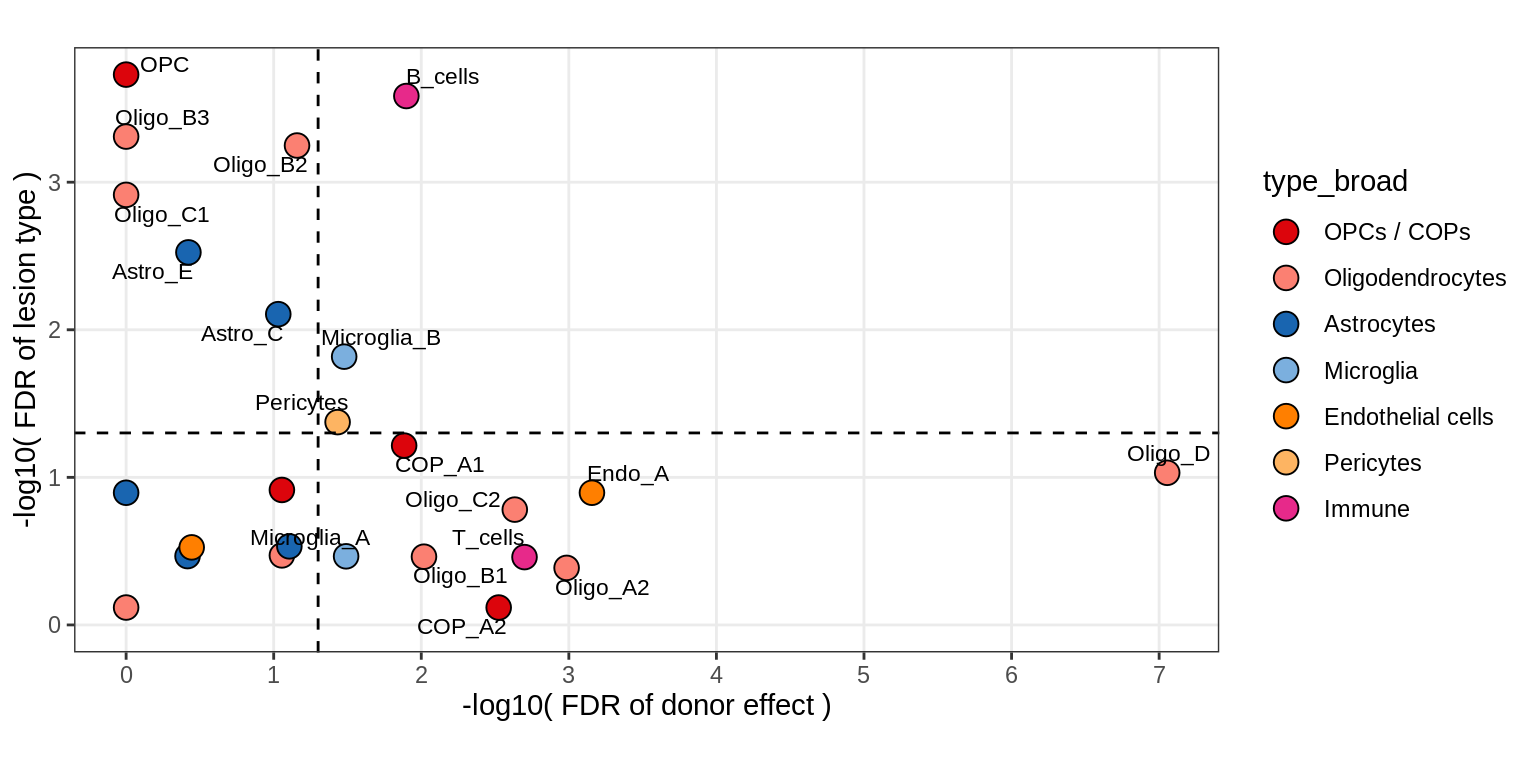
C
Contribution to variability in celltype abundances explained by lesion + patient in GM, no layers.
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_lrt_results-2.png", error = FALSE)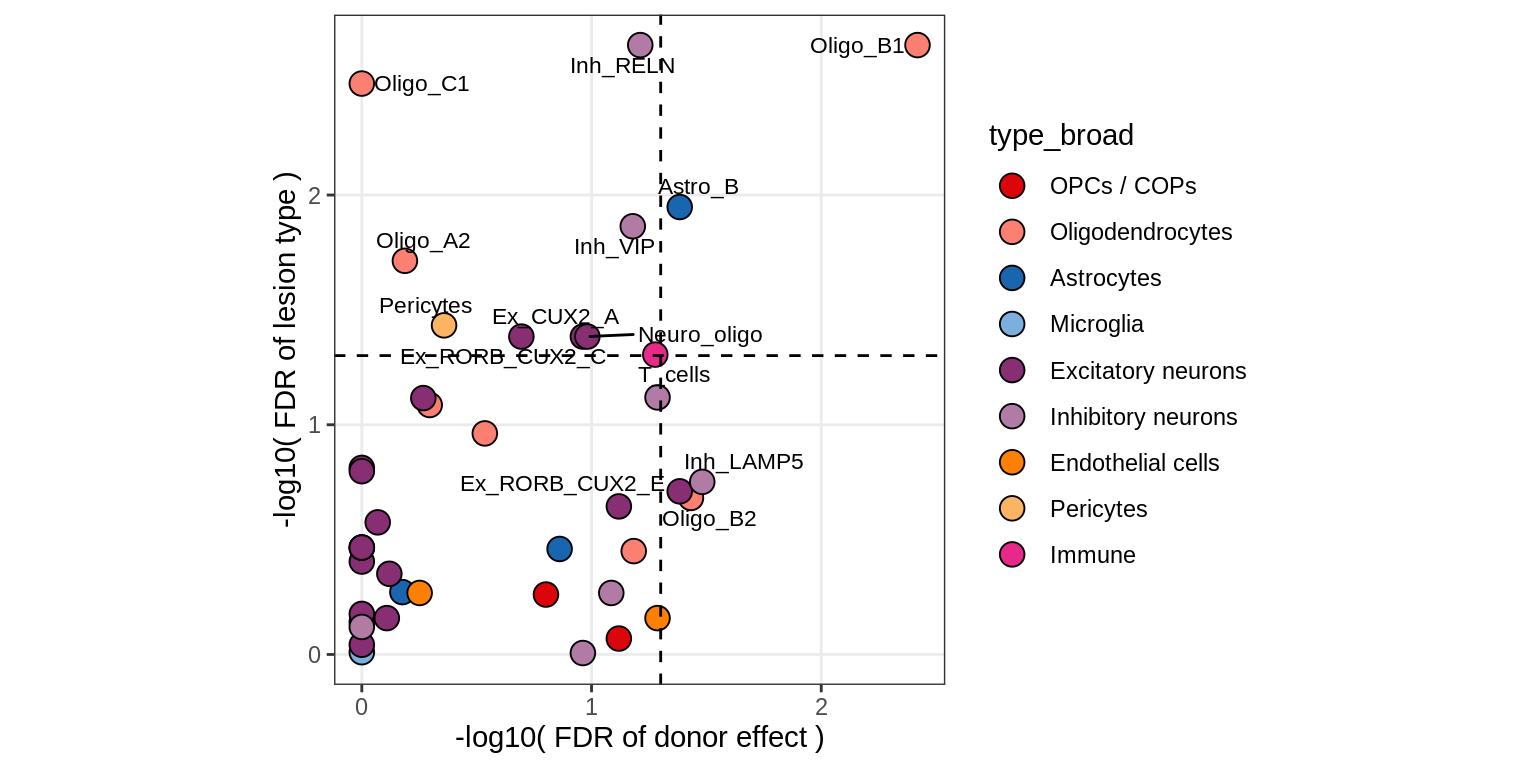
D
Differential abundance results for WM.
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_bootstraps_lesions_signif-1.png", error = FALSE)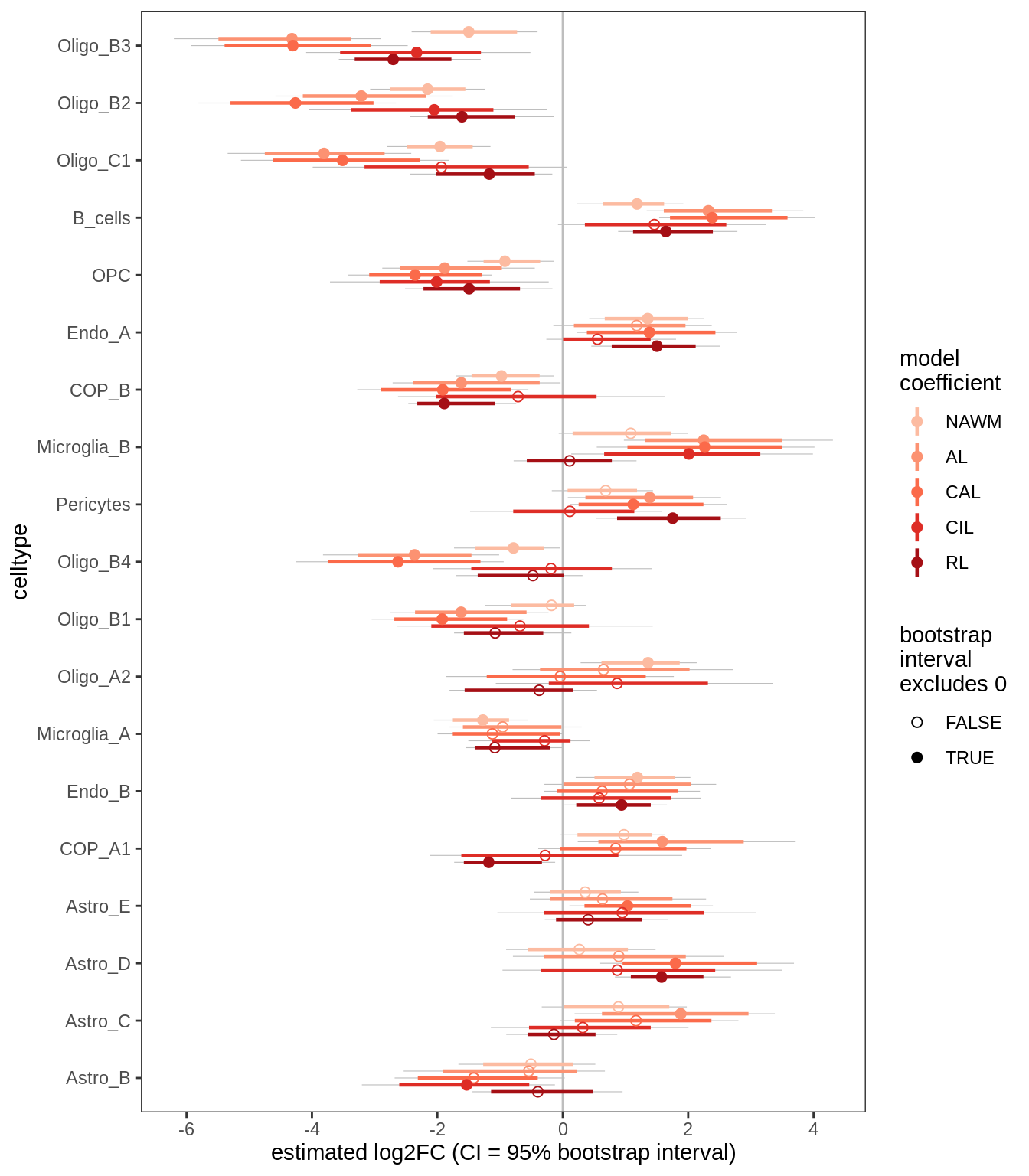
| Version | Author | Date |
|---|---|---|
| 270e5fc | wmacnair | 2021-11-26 |
E
Differential abundance results for GM (with layers factored out).
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_bootstraps_lesions_signif-6.png", error = FALSE)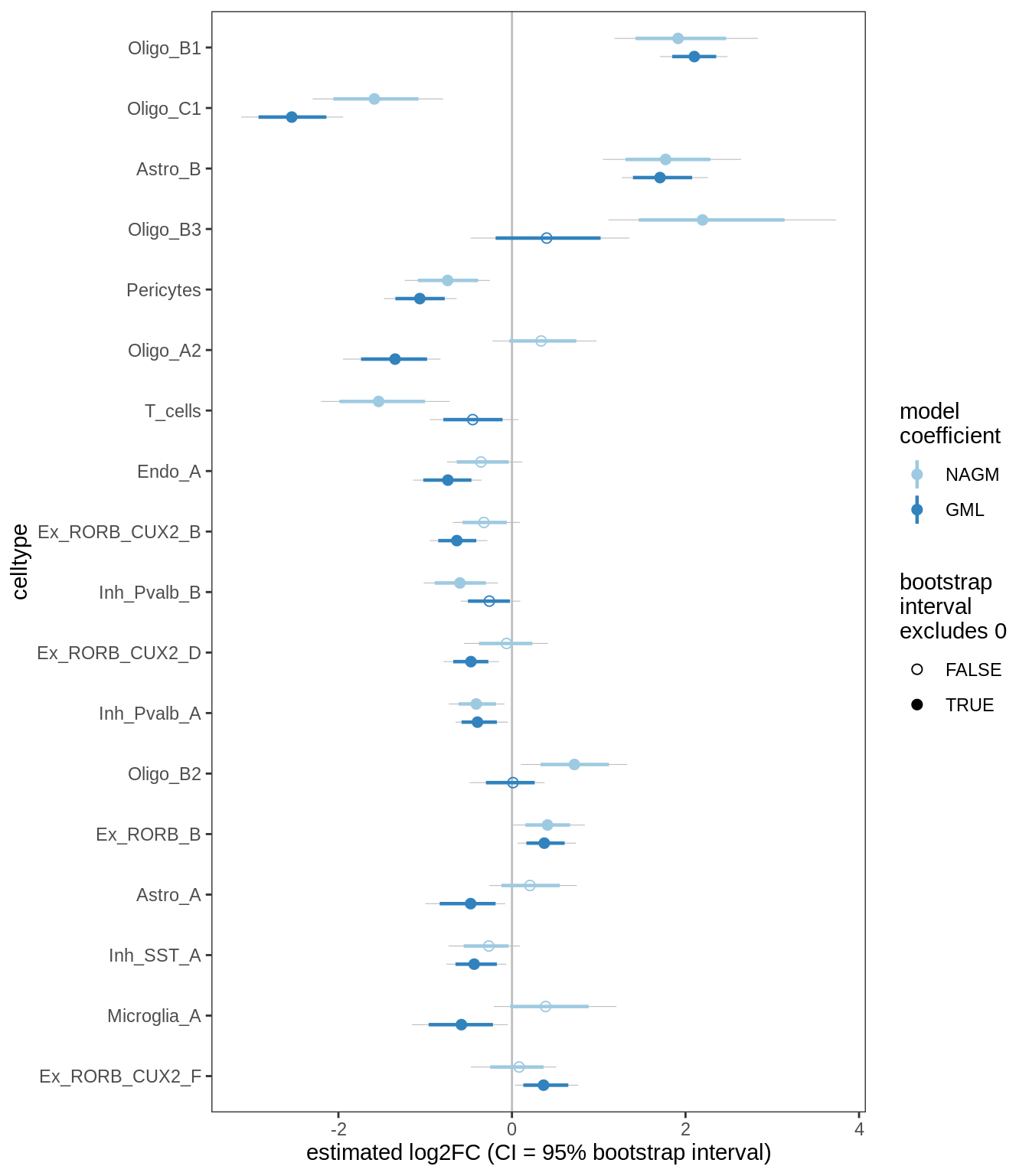
| Version | Author | Date |
|---|---|---|
| 270e5fc | wmacnair | 2021-11-26 |
Figure 3
A
include_graphics("figure/ms99_deg_figures_gm.Rmd/plot_de_barplot_sel-1.png", error = FALSE)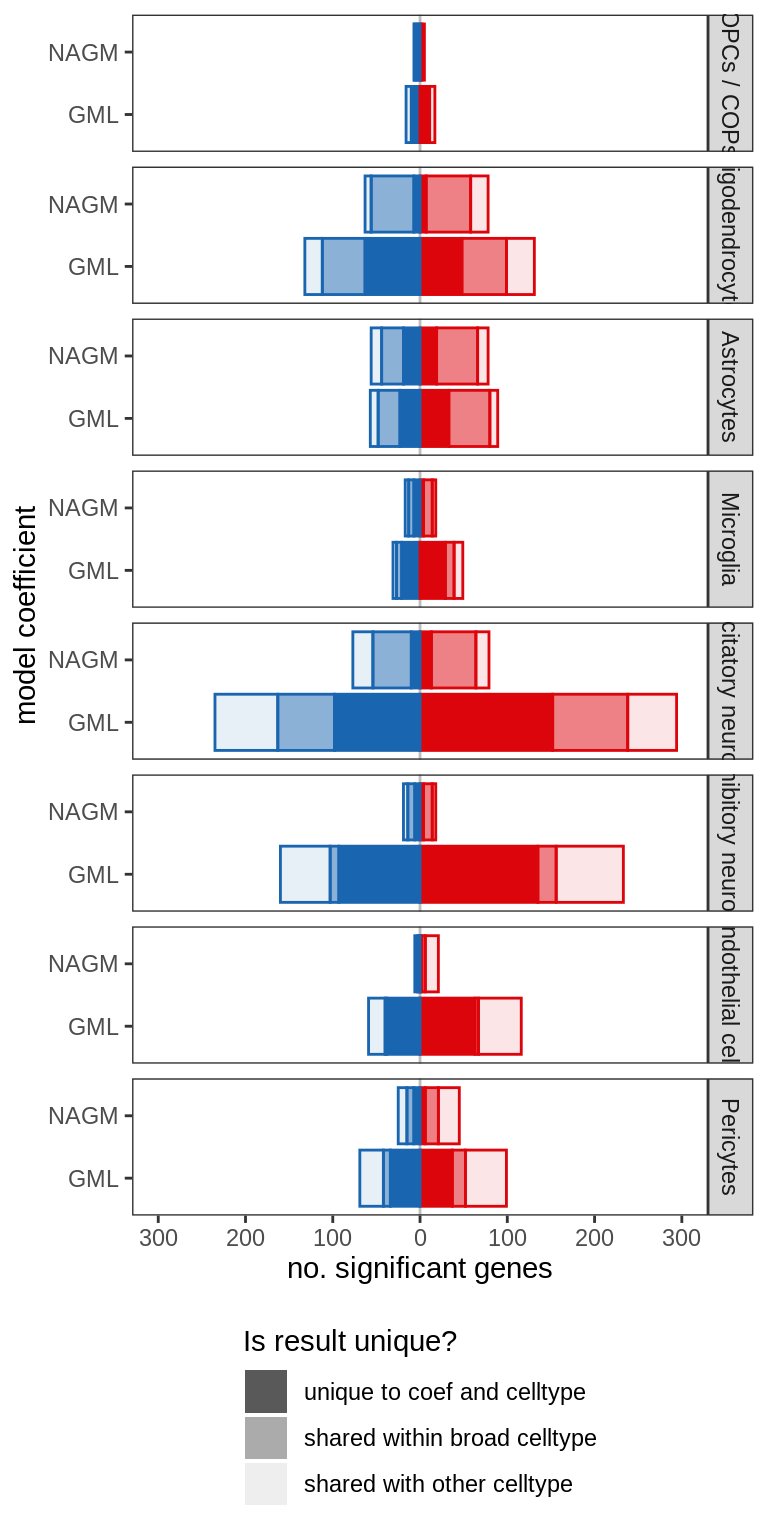
| Version | Author | Date |
|---|---|---|
| d83826d | wmacnair | 2022-01-27 |
B
include_graphics("figure/ms99_deg_figures_wm.Rmd/plot_de_barplot_sel-1.png", error = FALSE)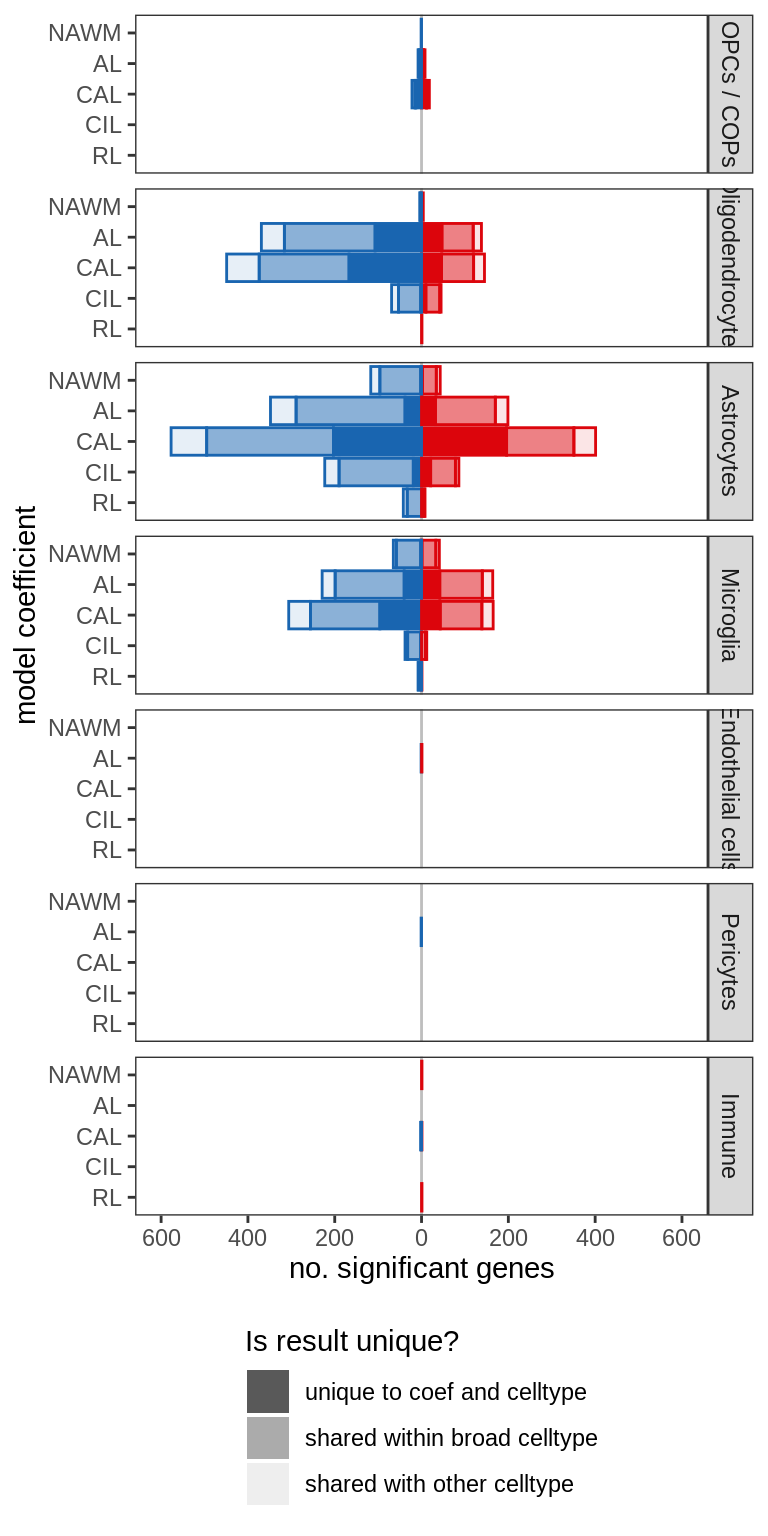
| Version | Author | Date |
|---|---|---|
| d83826d | wmacnair | 2022-01-27 |
C
Dotplot of Hallmark module results for GM.
[IN PRODUCTION]
D
Dotplot of Hallmark module results for WM.
[IN PRODUCTION]
E
Heatmap of selected interferon genes.
[IN PRODUCTION]
F
Genetic enrichment of differentially expressed genes.
include_graphics("figure/gwas_figures/de_barplots.png", error = FALSE)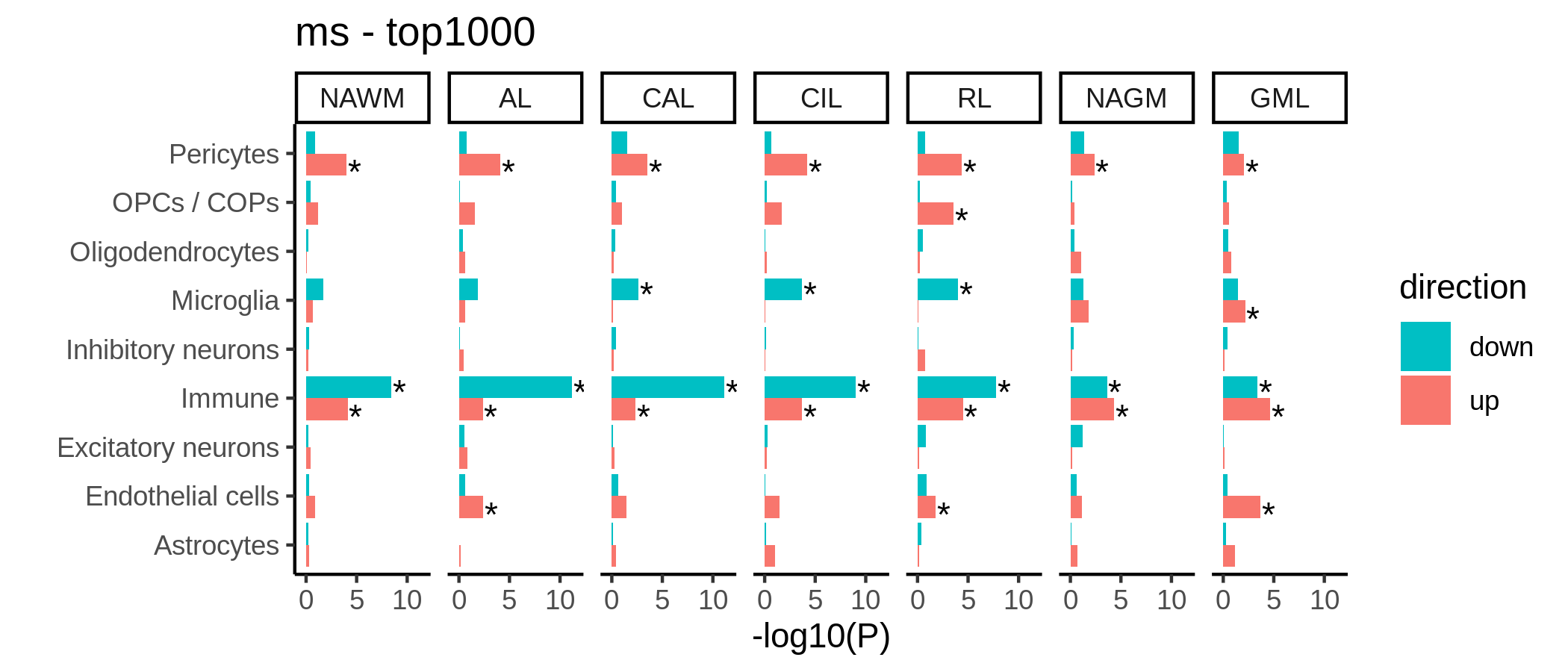
| Version | Author | Date |
|---|---|---|
| b1e52a7 | wmacnair | 2022-01-06 |
G
Clustering of WM fold change profiles. Restricted to genes where at least one lesion type has FDR < 5%. Clusters split so that average logFC difference between clusters is > log(4); clusters with fewer than 5 genes not shown; clusters ordered in descending order of mean logFC.
include_graphics("figure/ms99_deg_figures_wm.Rmd/plot_fc_cluster_profiles-1.png", error = FALSE)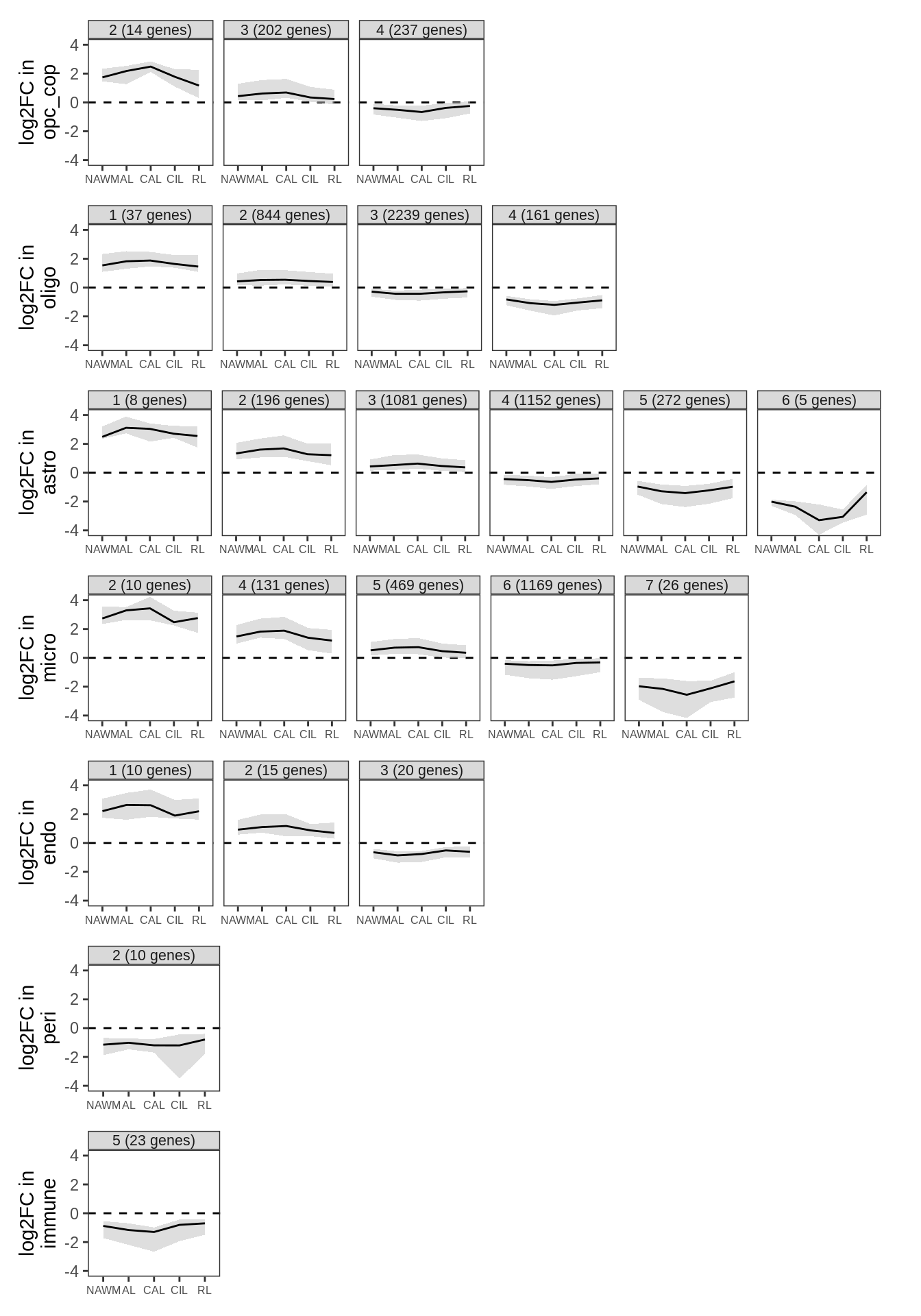
| Version | Author | Date |
|---|---|---|
| d83826d | wmacnair | 2022-01-27 |
Figure 4
A
Clustered expression heatmap of WM genes.
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/fig_overview_expression-2.png", error = FALSE)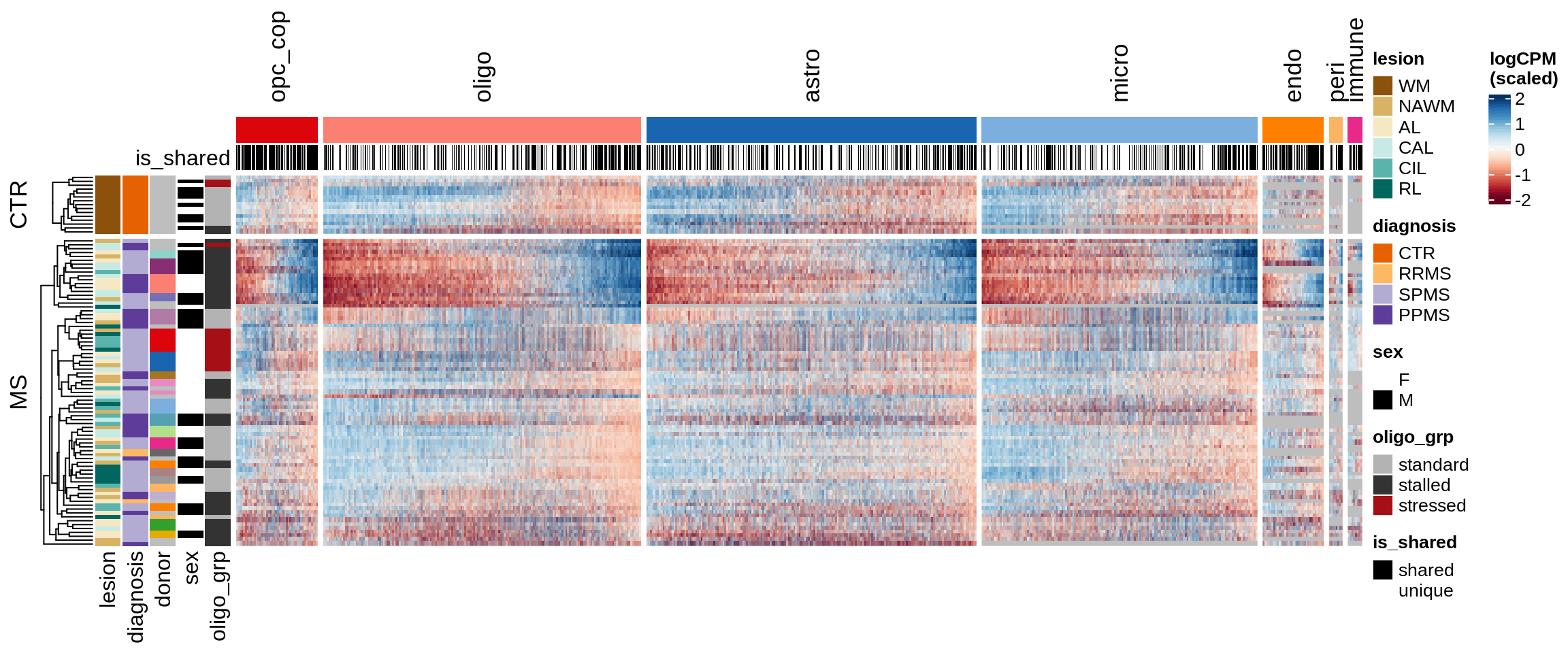
| Version | Author | Date |
|---|---|---|
| 8bc5188 | wmacnair | 2022-01-27 |
B
Variance explained
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/fig_factor_r2s-1.png", error = FALSE)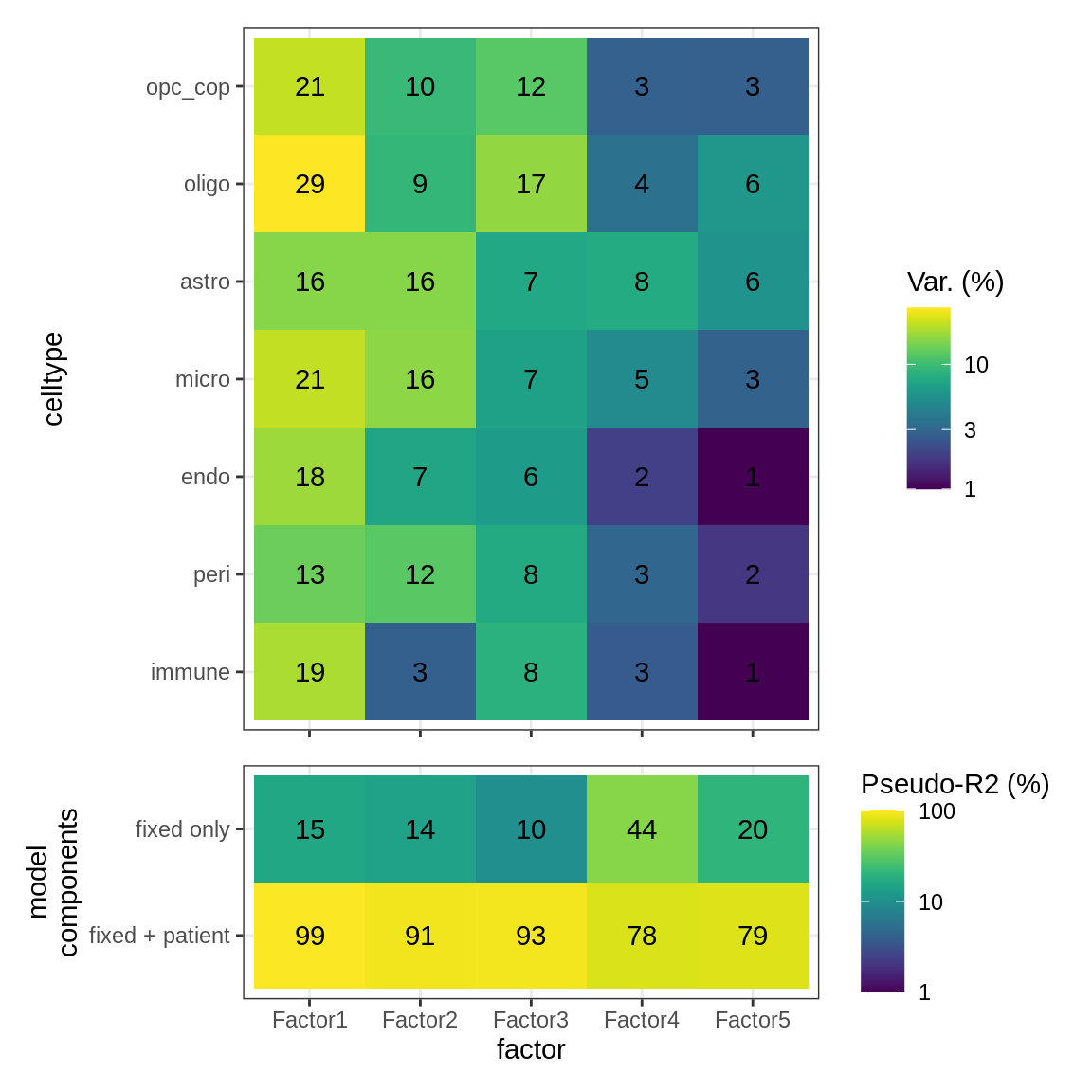
| Version | Author | Date |
|---|---|---|
| 8bc5188 | wmacnair | 2022-01-27 |
C
First two PCs of CLRs of oligodendroglia proportions.
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_sample_splits_clrs_oligos-6.png", error = FALSE)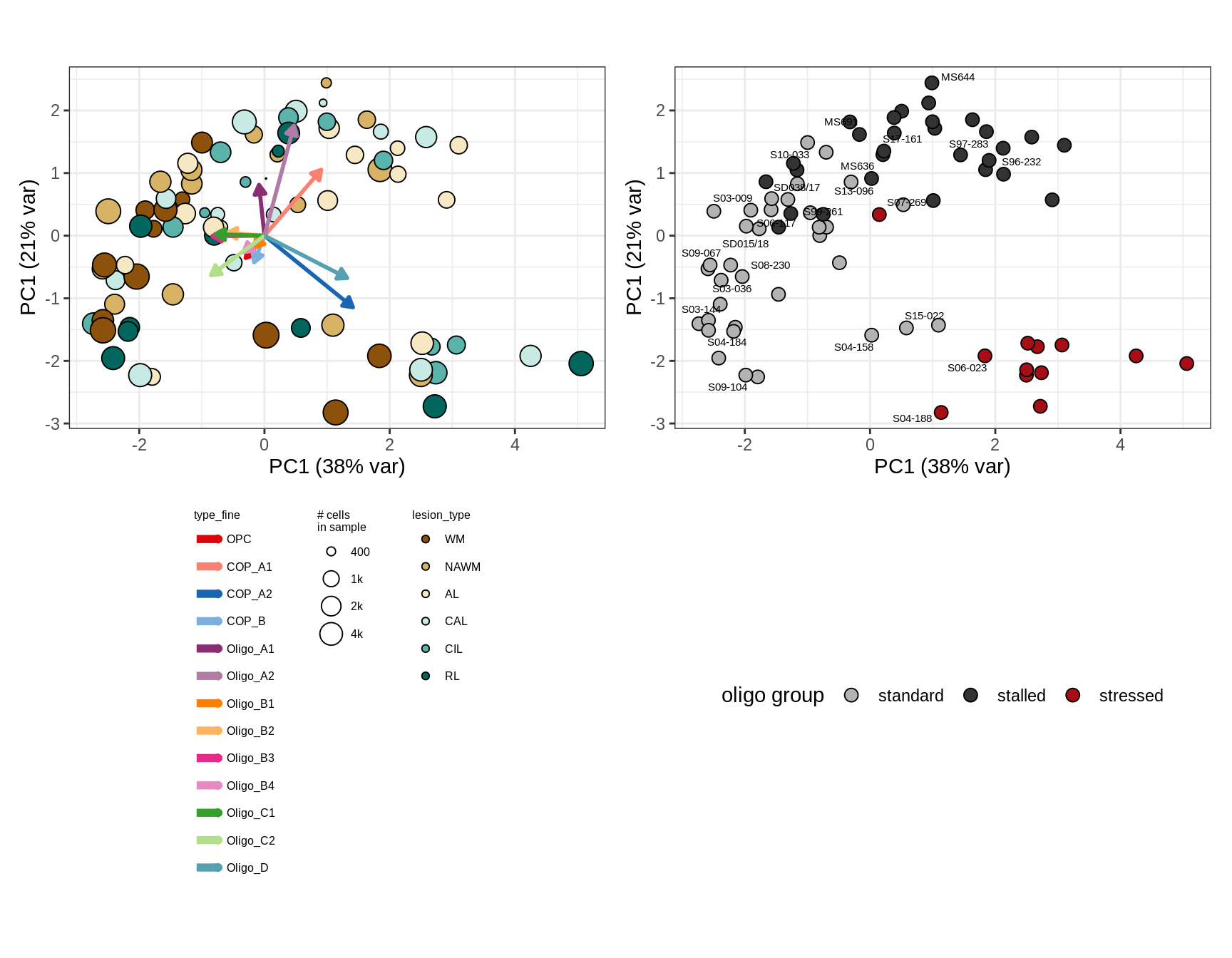
| Version | Author | Date |
|---|---|---|
| 8364a6f | wmacnair | 2021-12-13 |
D
Patient stratification via MOFA factors.
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/plot_factors_heatmap_few-1.png", error = FALSE)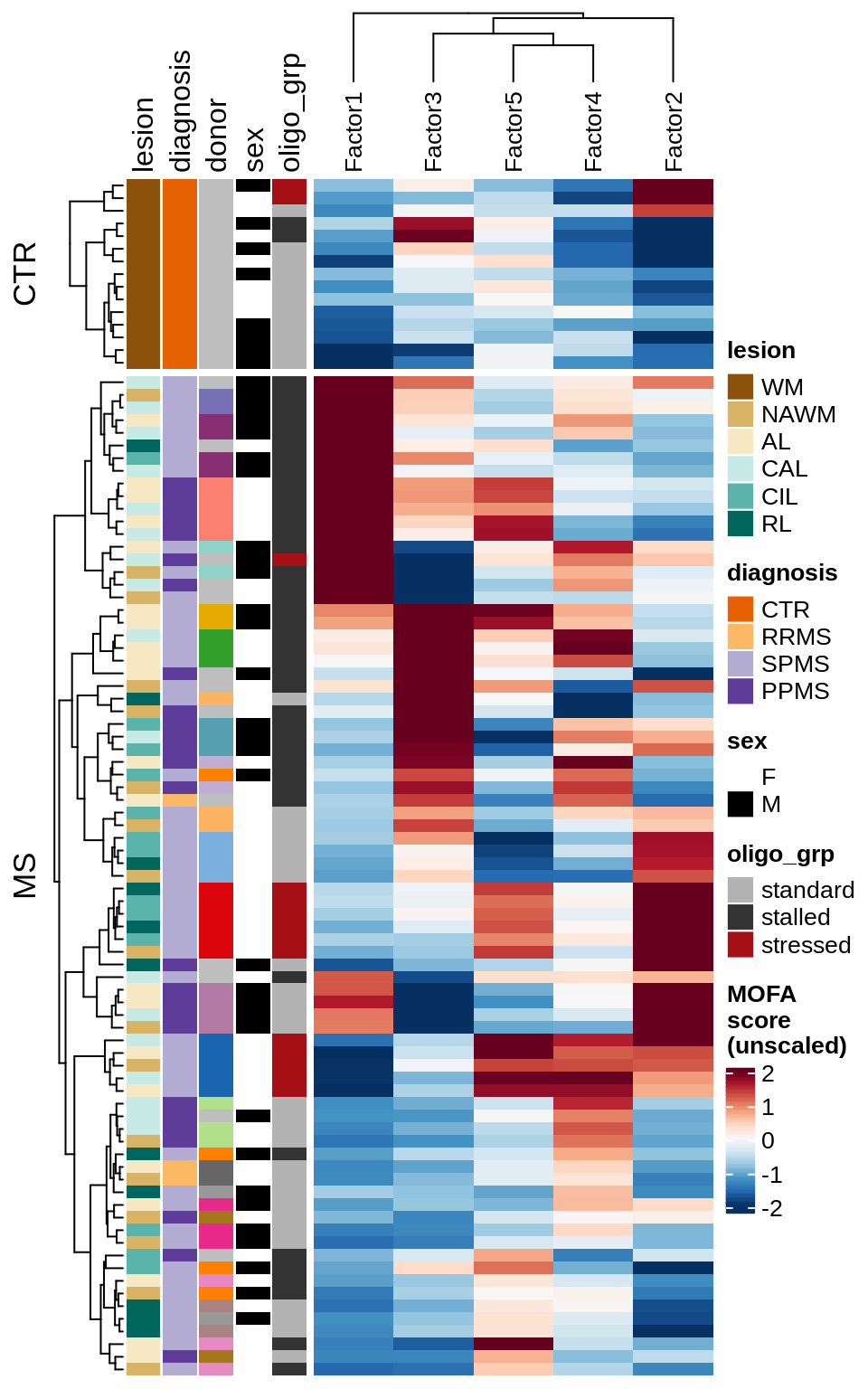
| Version | Author | Date |
|---|---|---|
| 8bc5188 | wmacnair | 2022-01-27 |
E
Factor 1 top genes
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/fig_factor1-1.png", error = FALSE)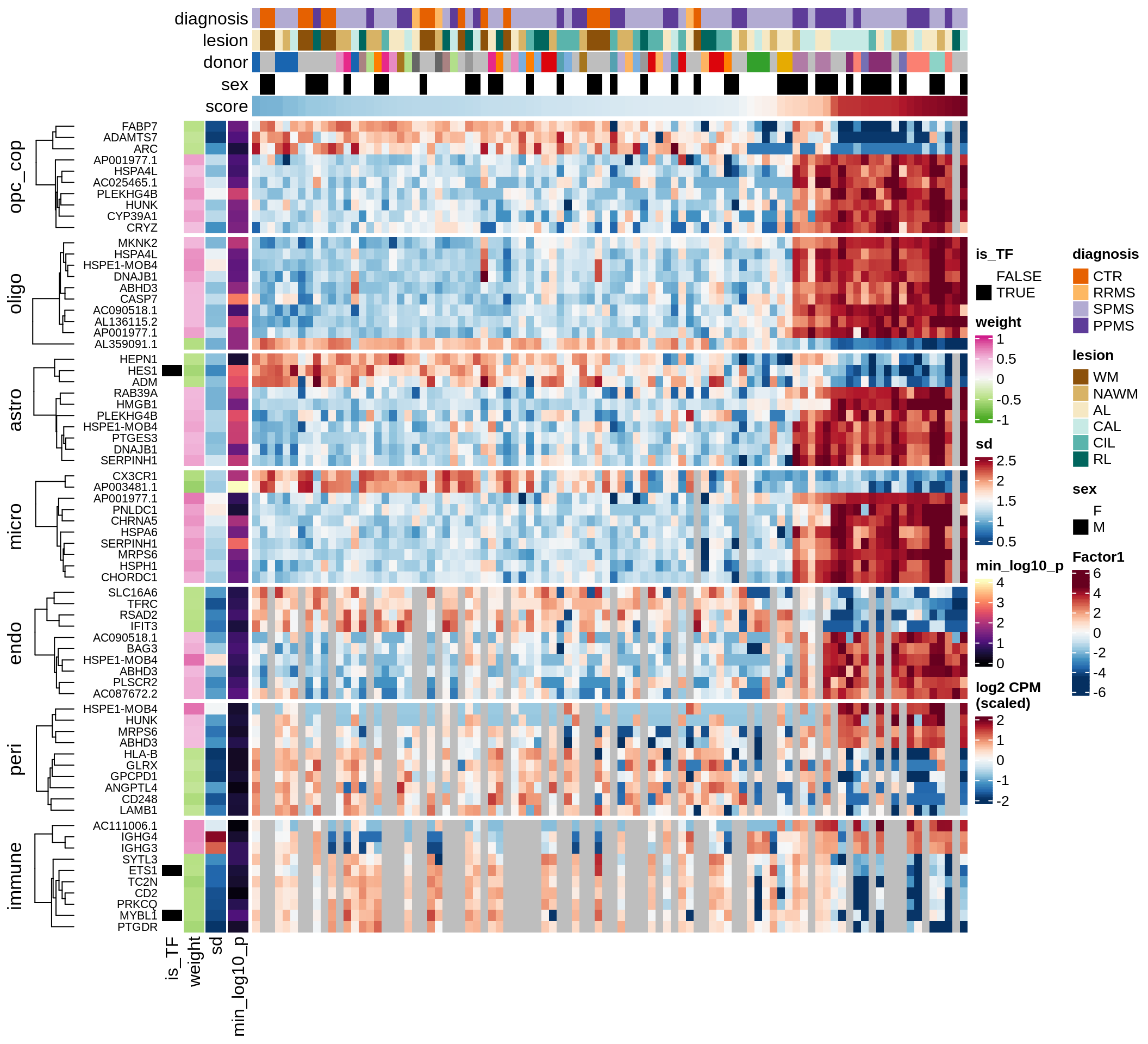
| Version | Author | Date |
|---|---|---|
| 8bc5188 | wmacnair | 2022-01-27 |
F
Factor 3 top genes
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/fig_factor3-1.png", error = FALSE)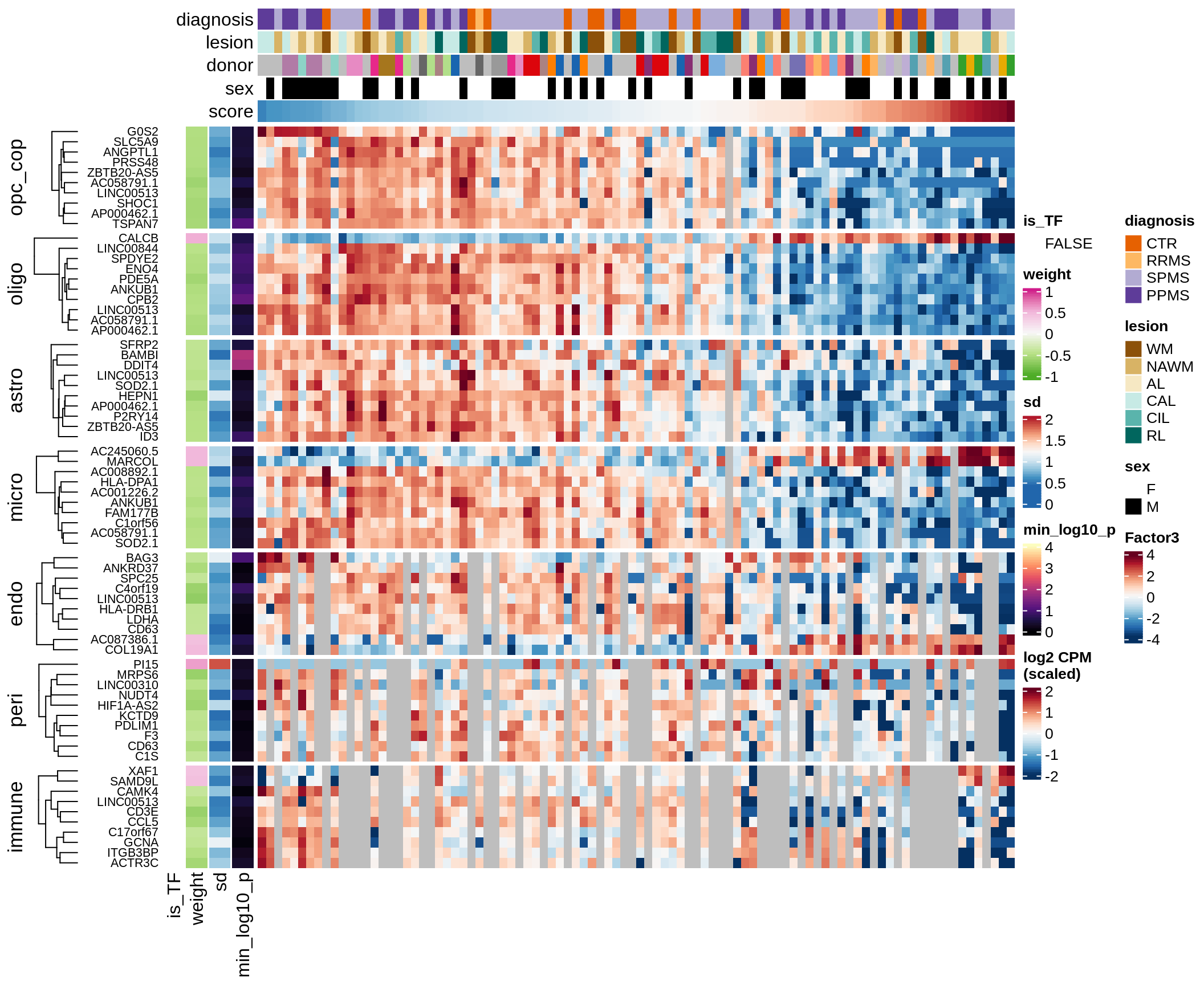
| Version | Author | Date |
|---|---|---|
| 8bc5188 | wmacnair | 2022-01-27 |
G
Factor 5 top genes
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/fig_factor5-1.png", error = FALSE)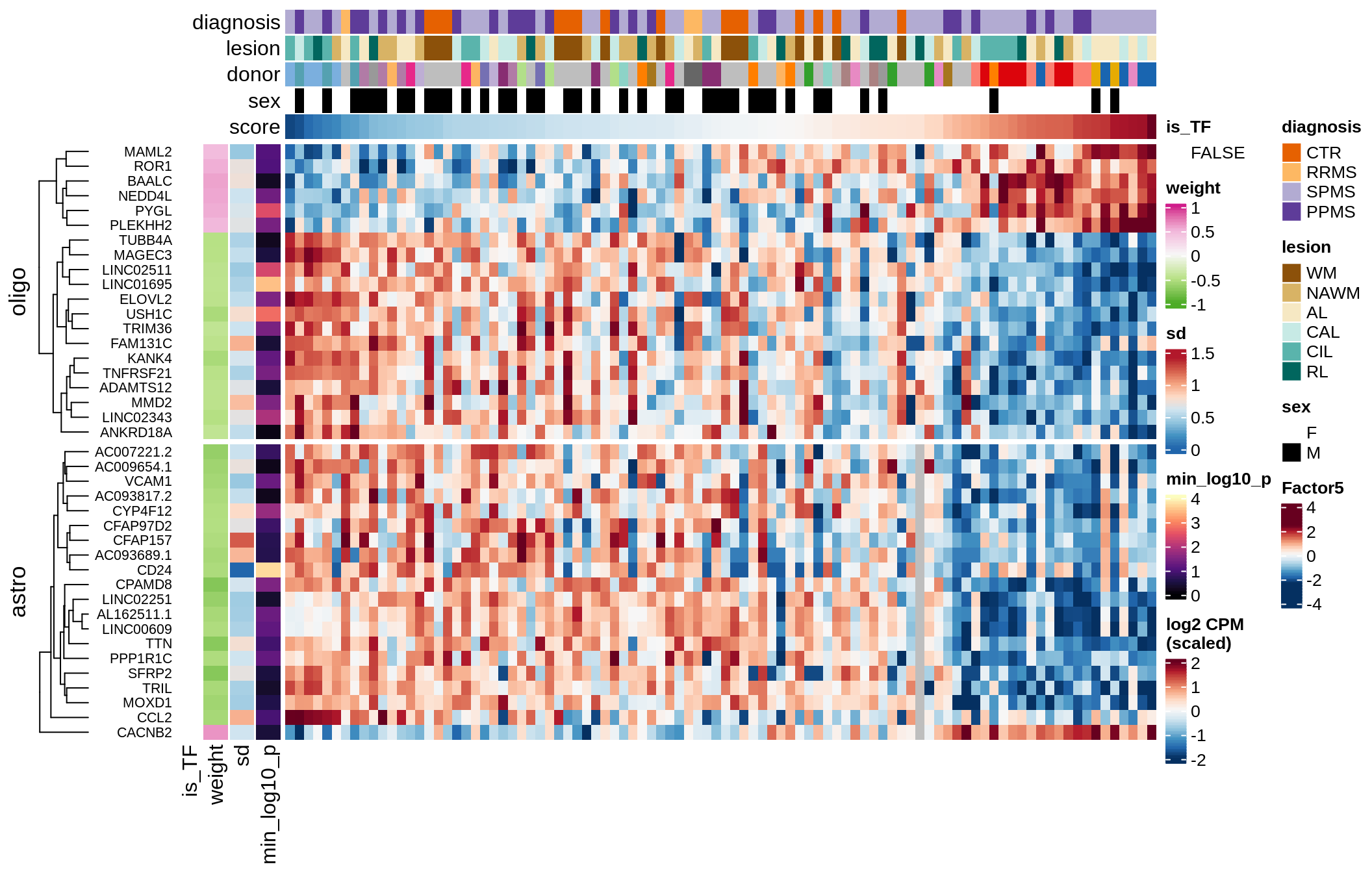
| Version | Author | Date |
|---|---|---|
| 8bc5188 | wmacnair | 2022-01-27 |
Supplementary figures
Unused figure / supp figs I haven’t sorted out yet
Sx
Post-QC summary of samples
include_graphics("de_reports/figure/ms03_SampleQC.Rmd/plot_totals_split_by_meta-1.png", error = FALSE)
Sx
[SCCAF]
include_graphics("de_reports/figure/ms03_SampleQC.Rmd/plot_totals_split_by_meta-1.png", error = FALSE)Sx
Expression of marker genes selected for broad celltypes, and for fine celltypes. Expression calculated across all cells and samples.
include_graphics("figure/ms12_markers.Rmd/plot_dotplot_dheeraj_compact-1.png", error = FALSE)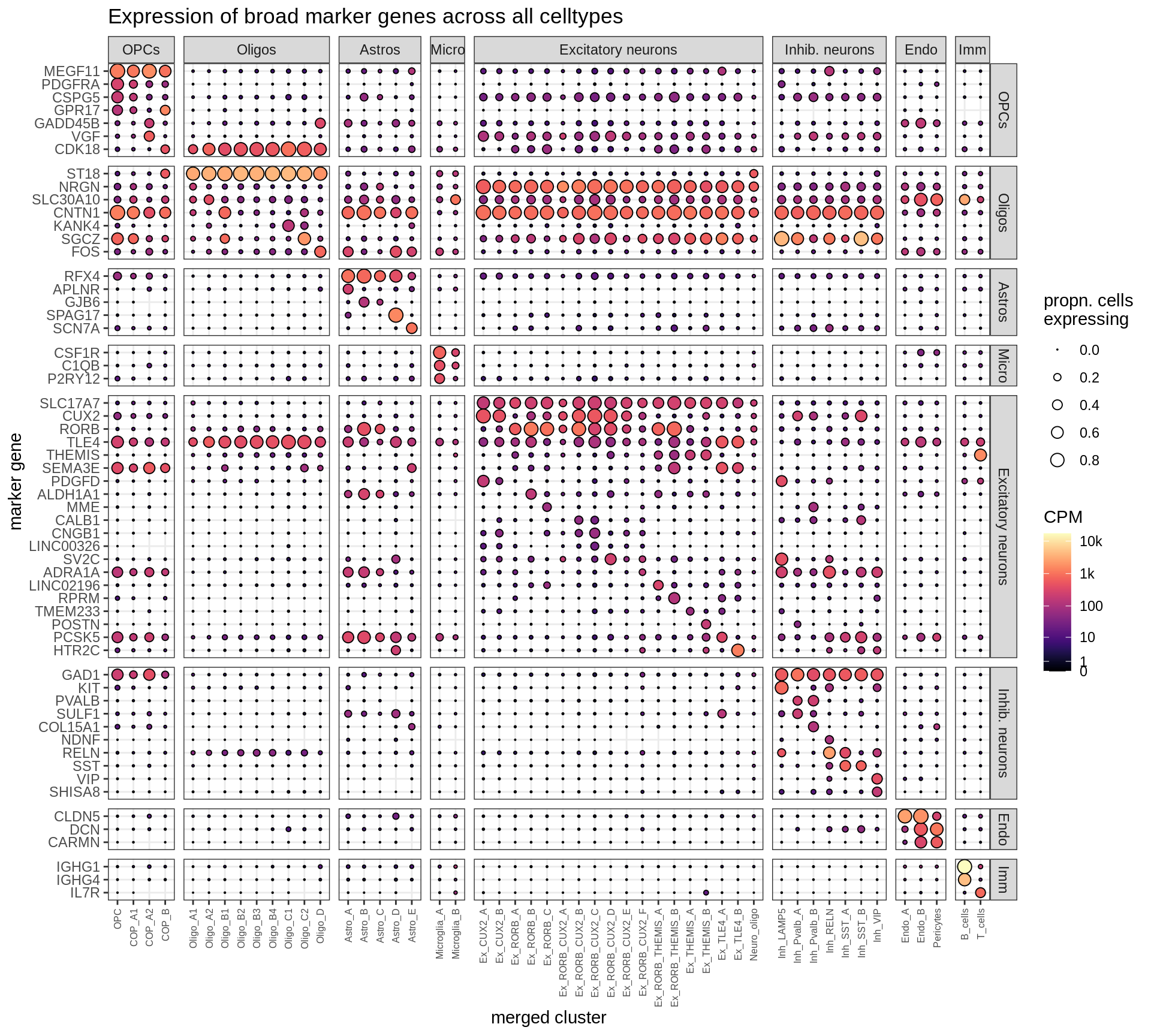
| Version | Author | Date |
|---|---|---|
| 7fb1b95 | wmacnair | 2021-11-25 |
Sx
[Cluster mixing]
include_graphics("figure/ms04_conos.Rmd/plot_conos_mixing-1.png", error = FALSE)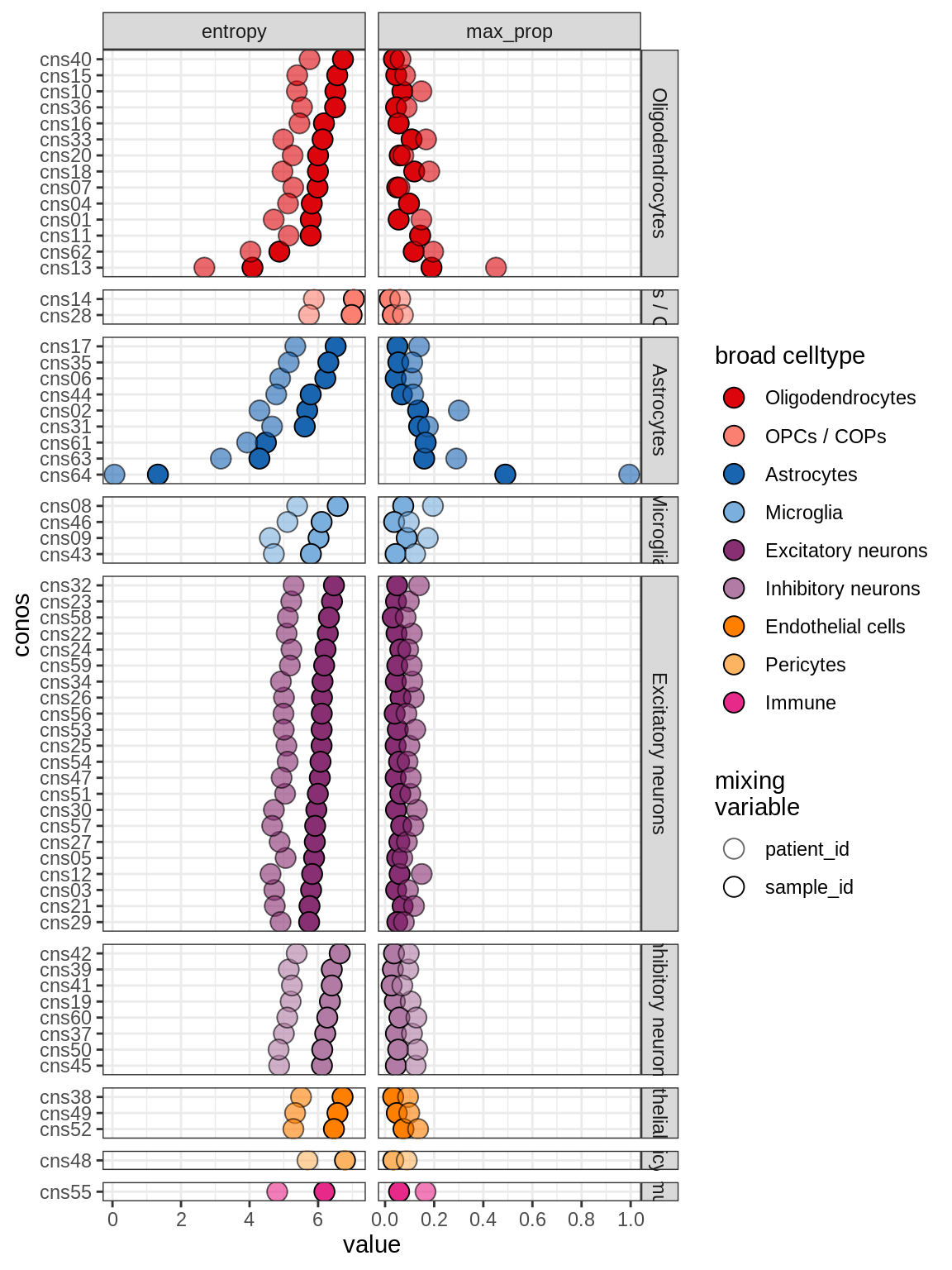
| Version | Author | Date |
|---|---|---|
| 7fb1b95 | wmacnair | 2021-11-25 |
Sx
[GM layer effects]
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_propns_layers-1.png", error = FALSE)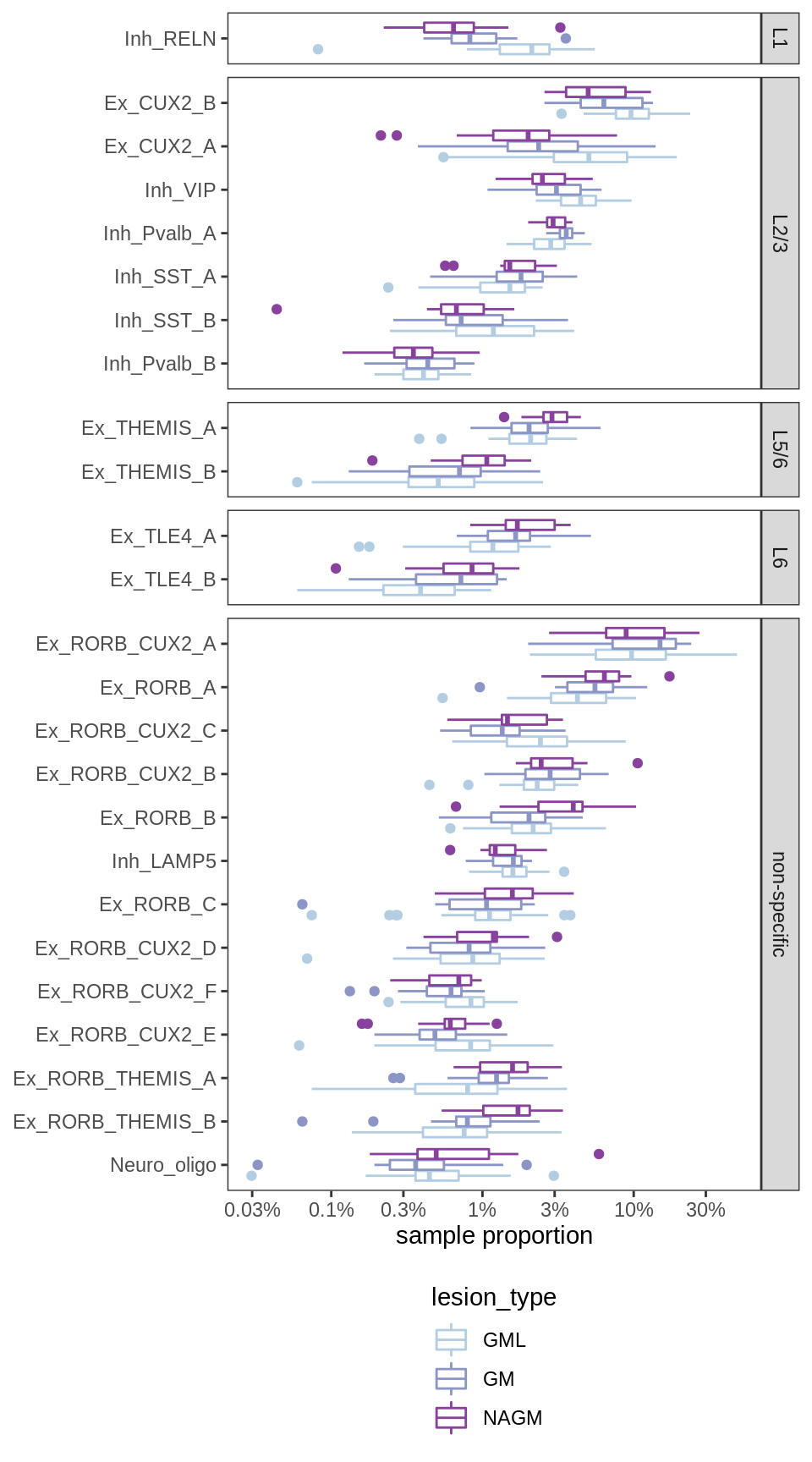
Sx
WM oligodendroglia proportions barplot
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_sample_splits_bars_oligos-1.png", error = FALSE)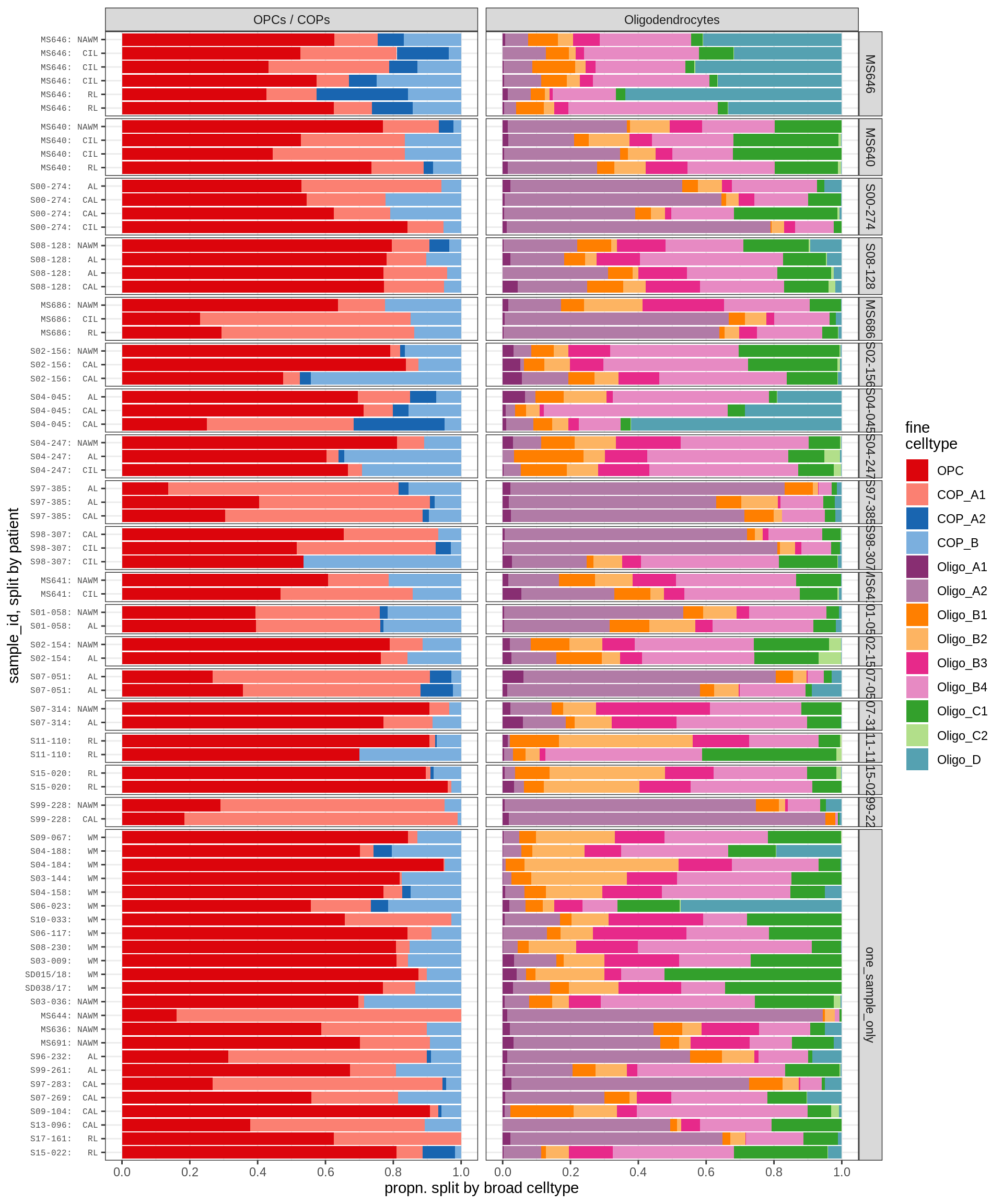
Sx
Comparison of Sarah’s validation of GPR17-expressing cells
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_no_gpr17_cells-1.png", error = FALSE)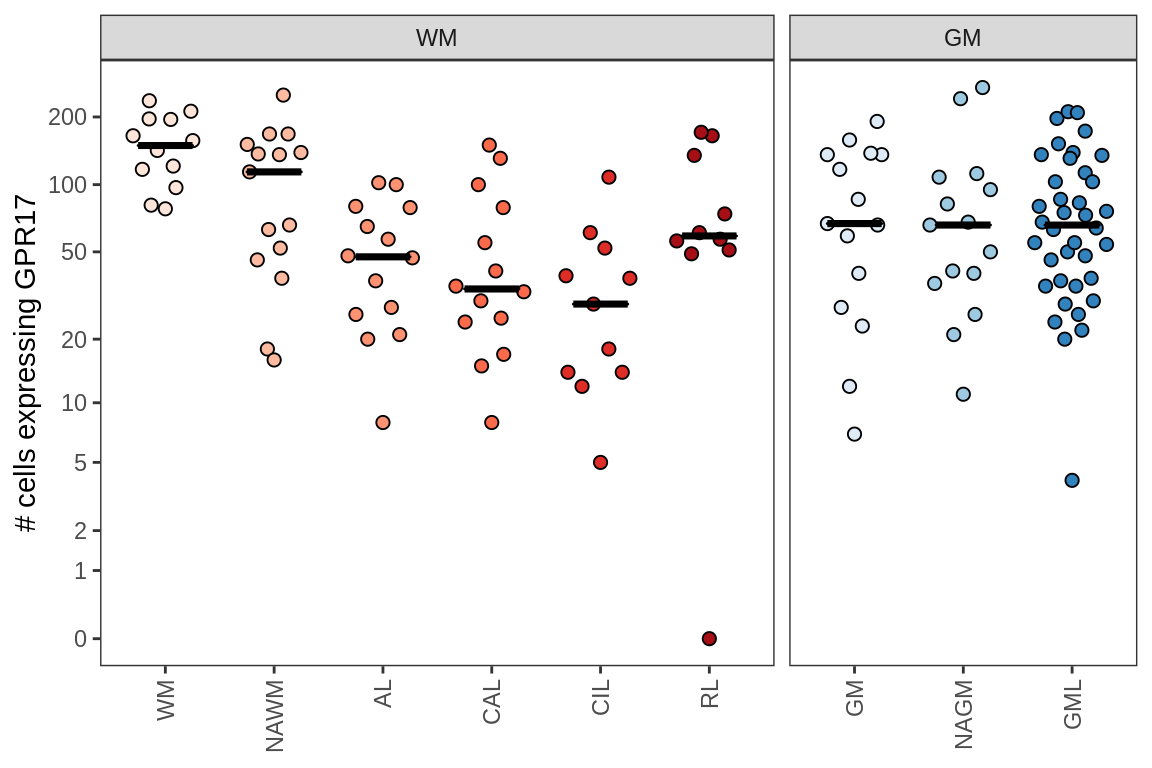
| Version | Author | Date |
|---|---|---|
| afba18d | wmacnair | 2021-12-20 |
Sx
Contribution to variability in celltype abundances explained by lesion + patient in GM, including 4 layer PCs
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_lrt_results-6.png", error = FALSE)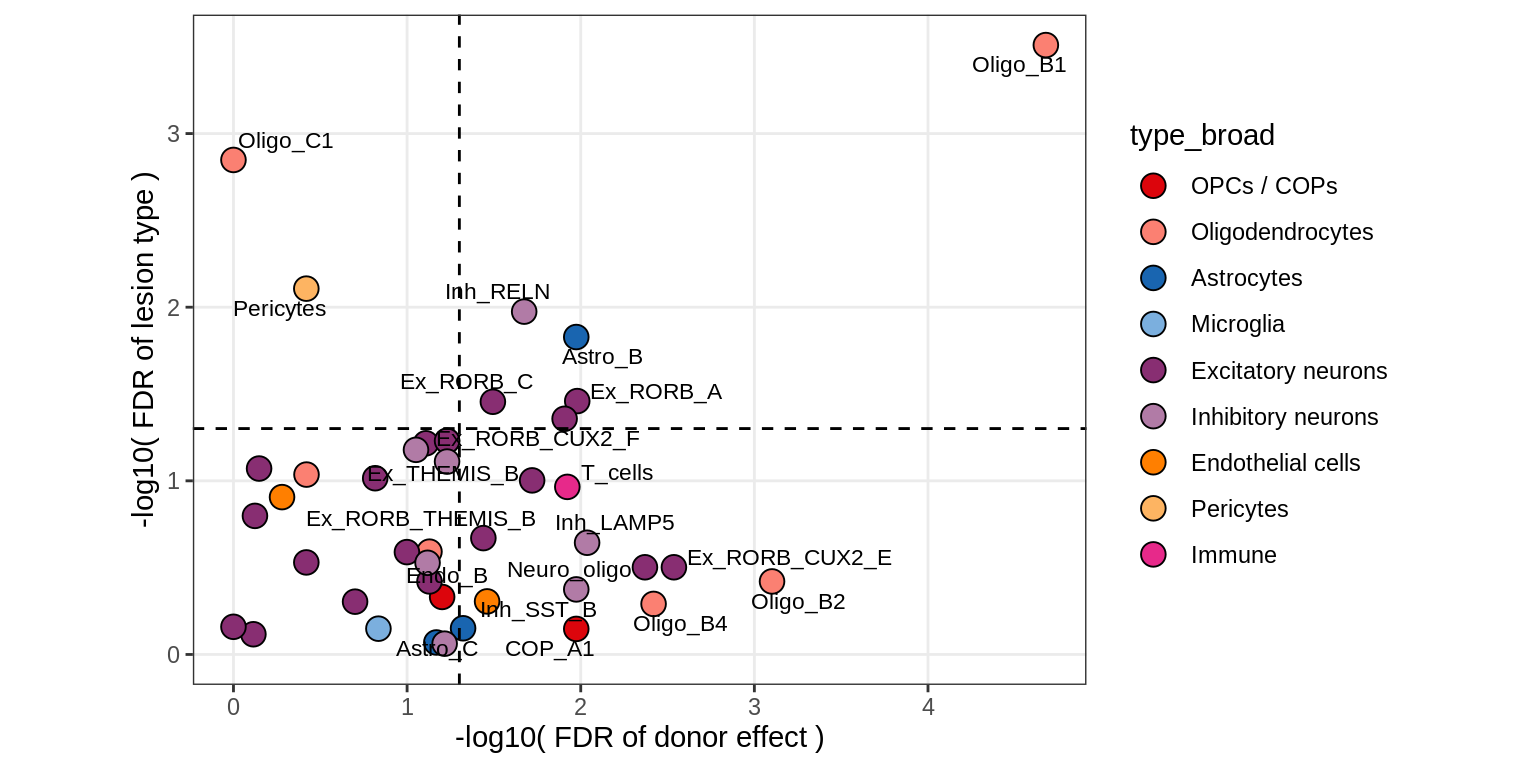
Sx
DA results for GM, no layers
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_bootstraps_lesions-2.png", error = FALSE)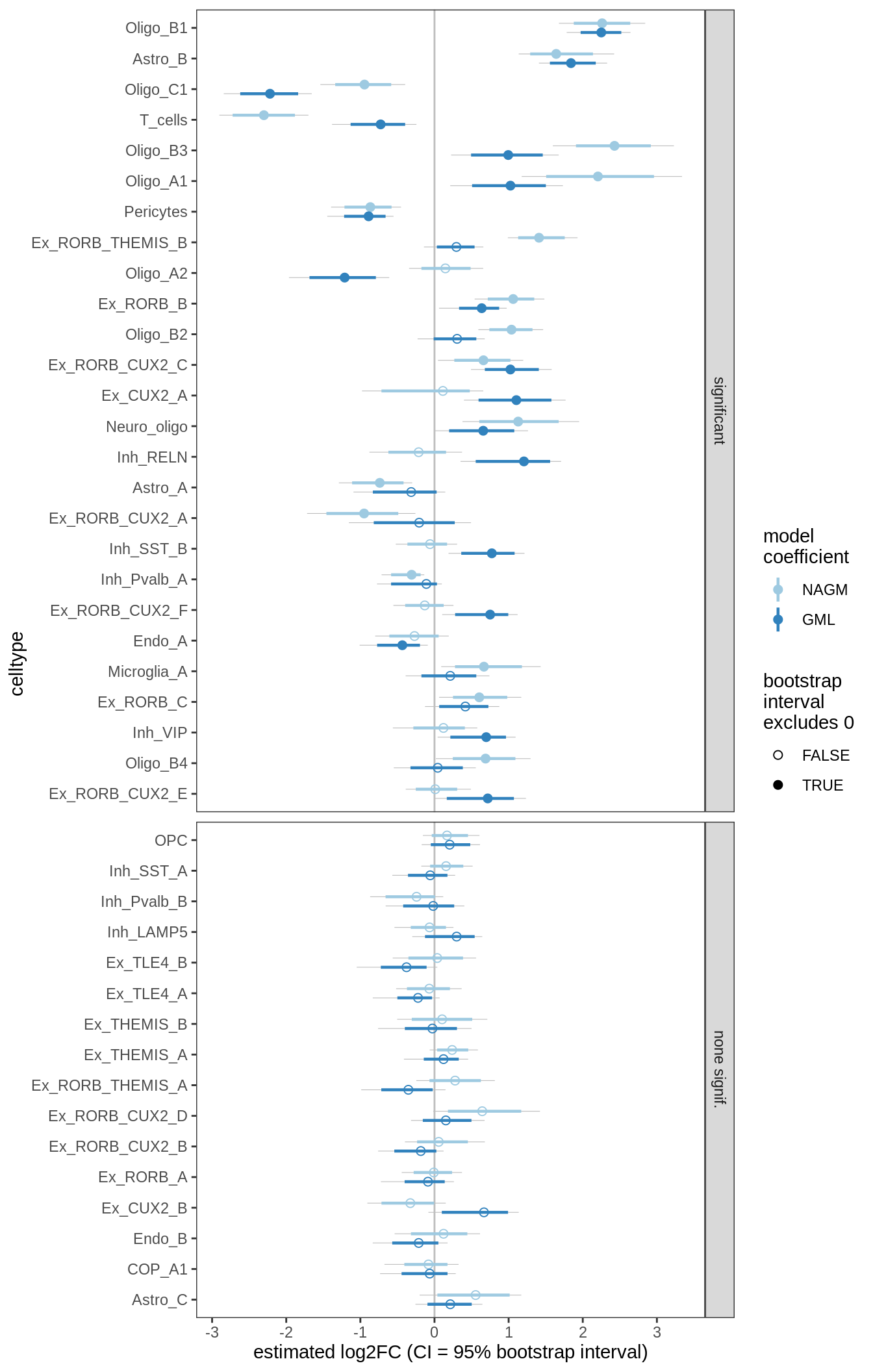
Sx
include_graphics("figure/ms99_deg_figures_wm.Rmd/plot_causes_of_variability-1.png", error = FALSE)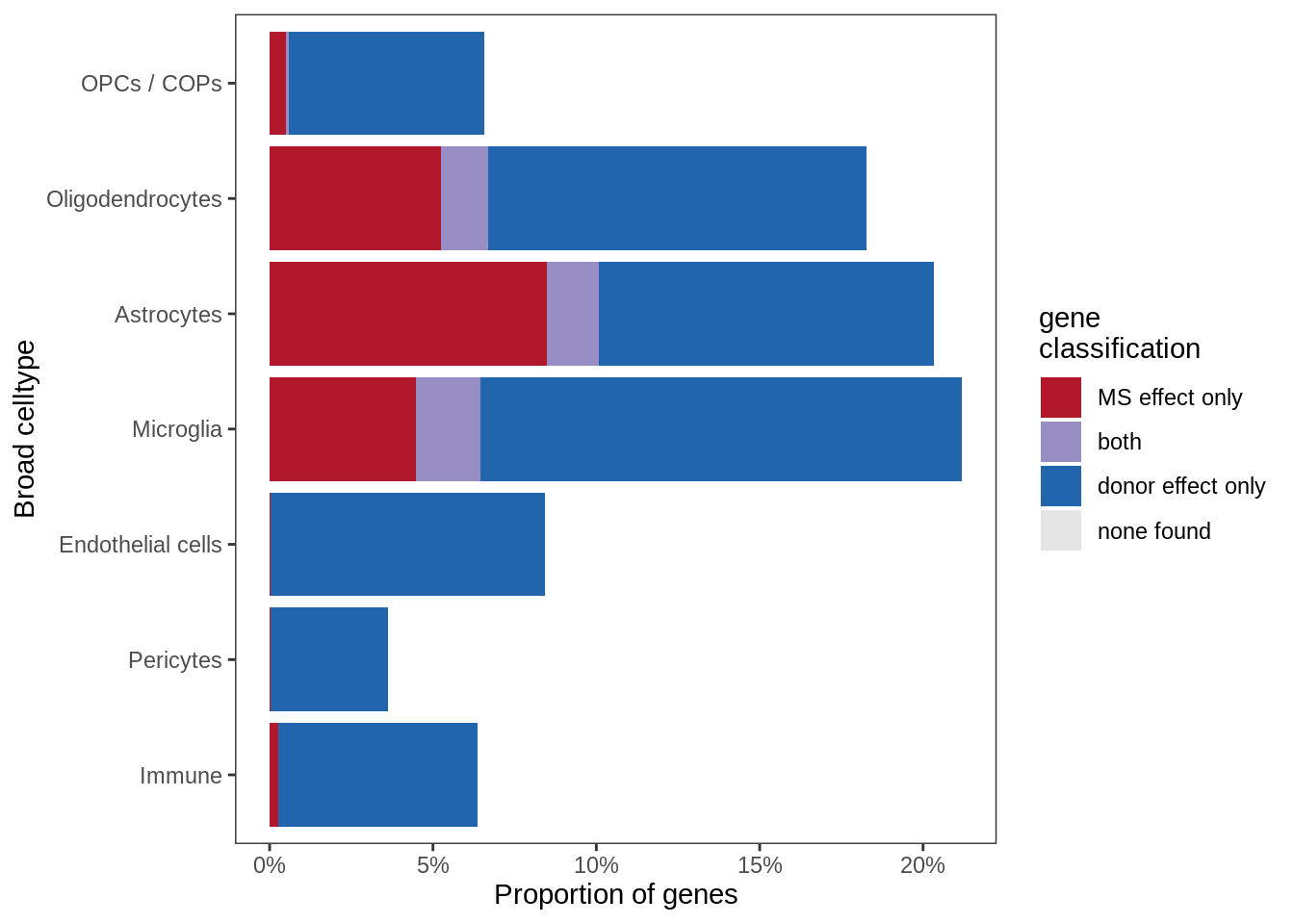
Sx
include_graphics("figure/ms99_deg_figures_gm.Rmd/plot_causes_of_variability-1.png", error = FALSE)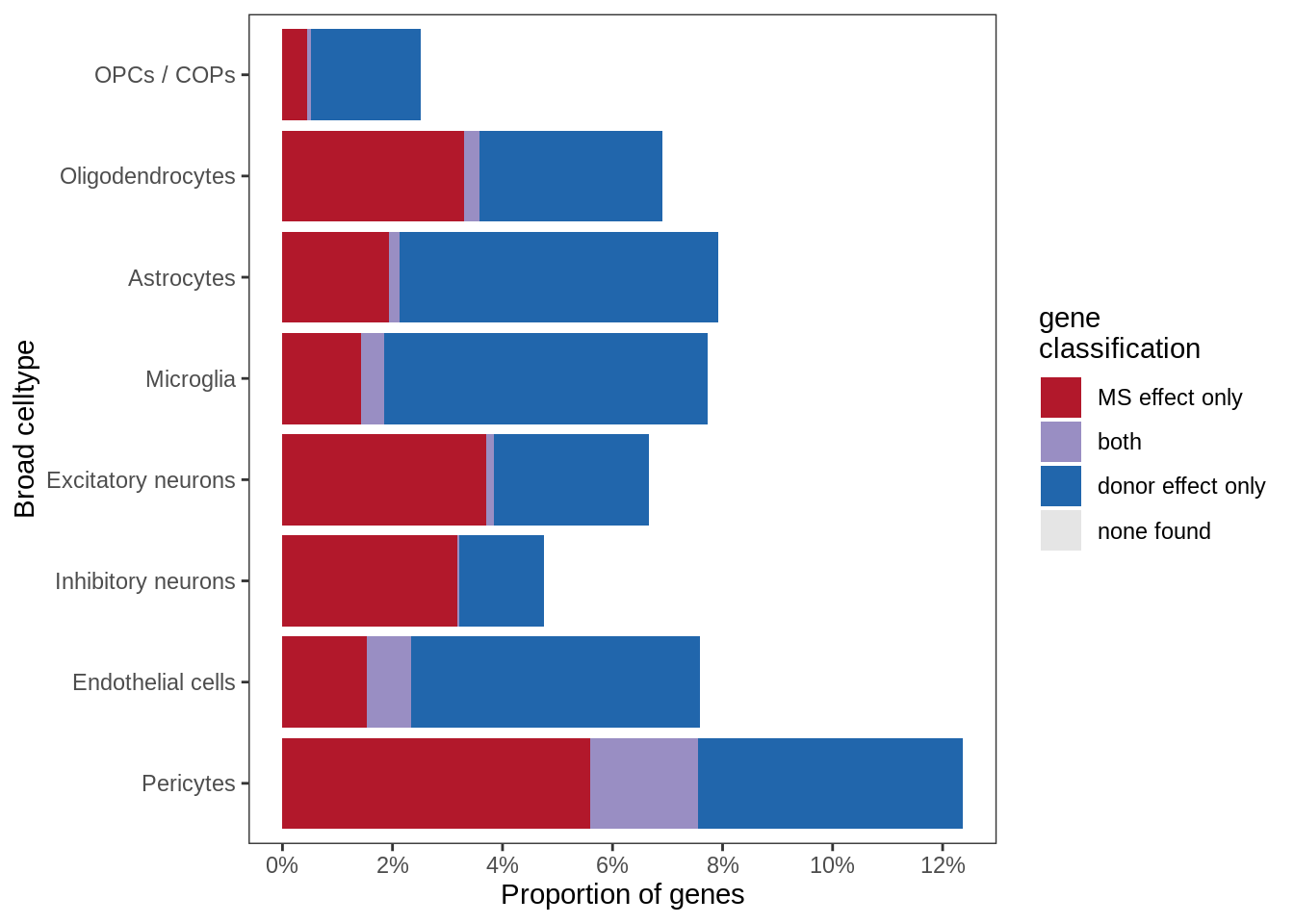
Sx
include_graphics("figure/ms15_mofa_sample_gm_w_layers_final_meta.Rmd/fig_overview_expression-2.png", error = FALSE)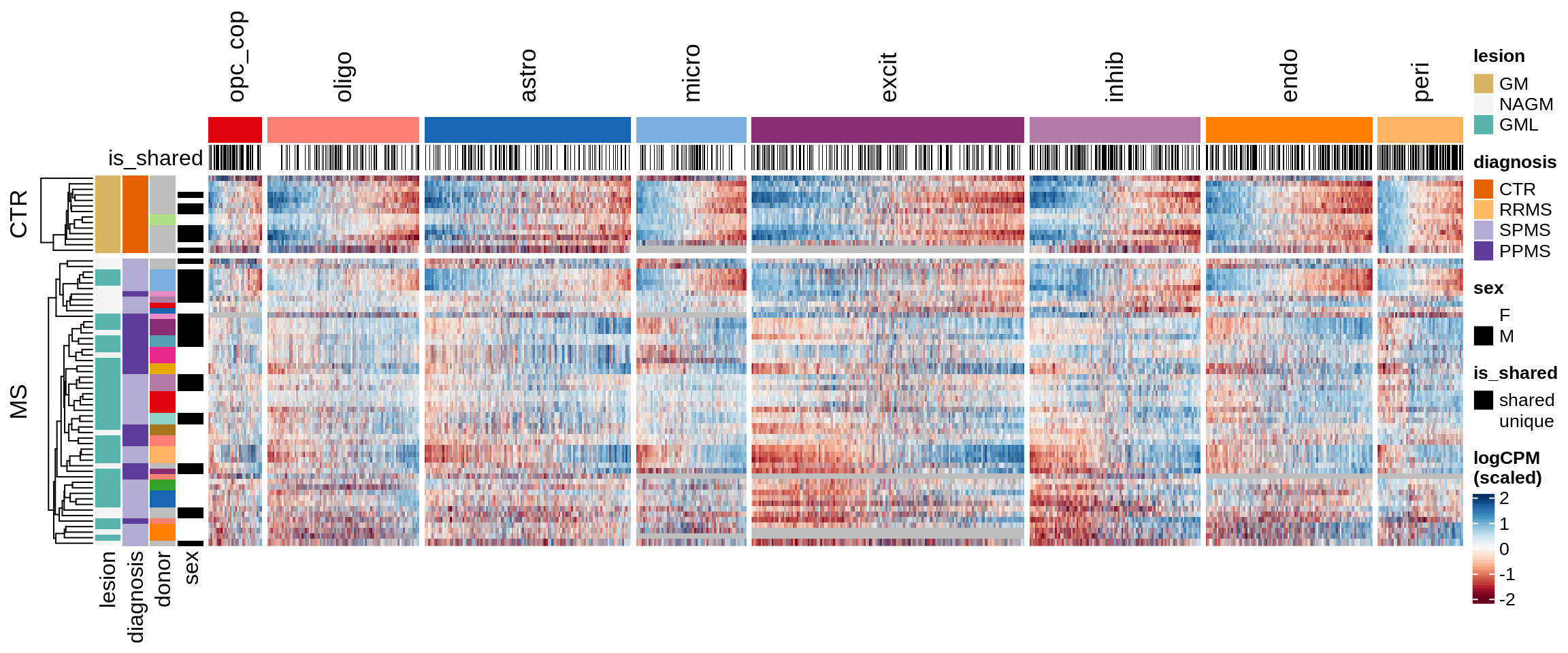
| Version | Author | Date |
|---|---|---|
| 1d0d7e8 | wmacnair | 2022-01-21 |
Sx
Effect of including PCs, neurons
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_effect_of_pcs_lesions-1.png", error = FALSE)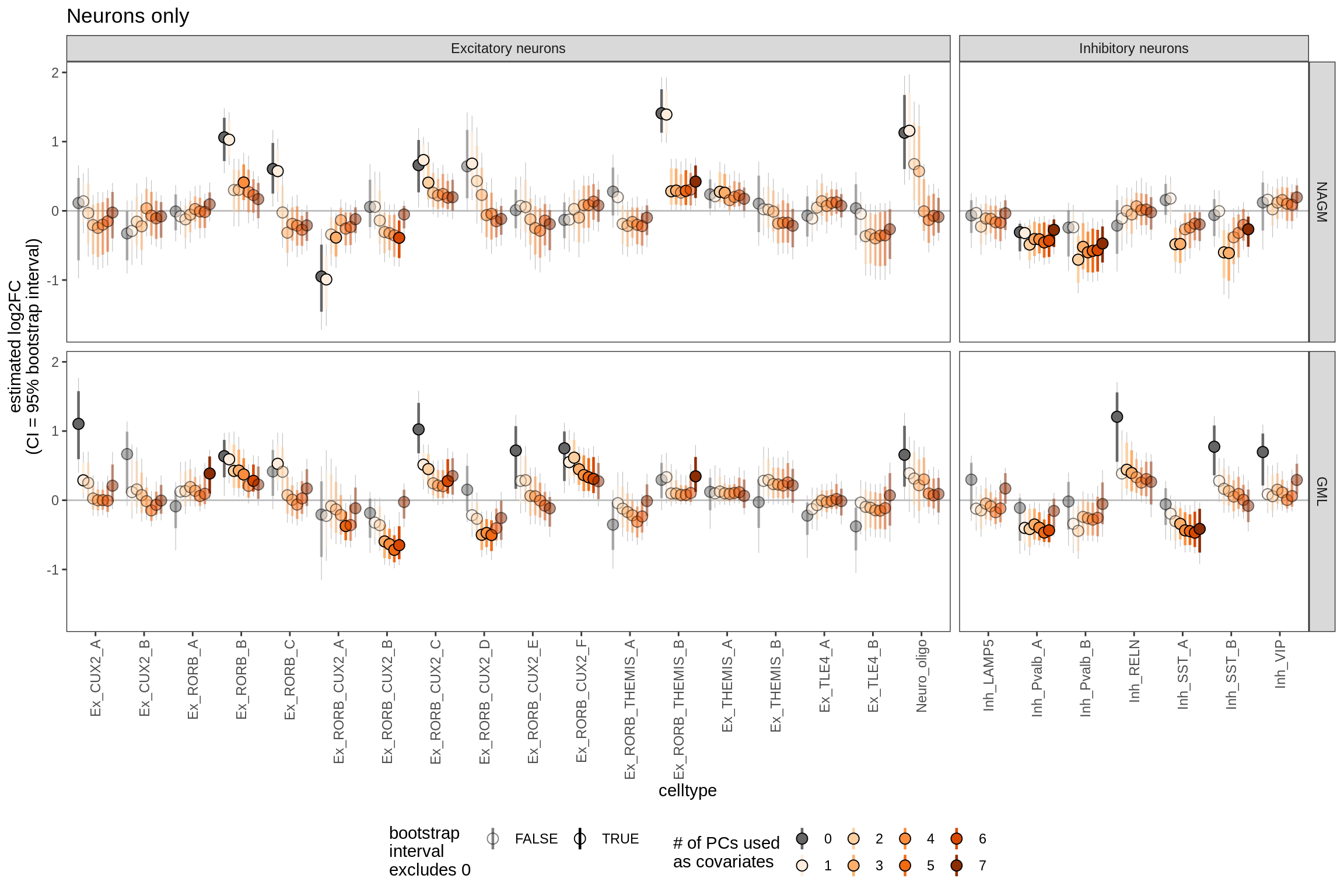
Sx
Effect of including PCs, other celltypes
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_effect_of_pcs_lesions-2.png", error = FALSE)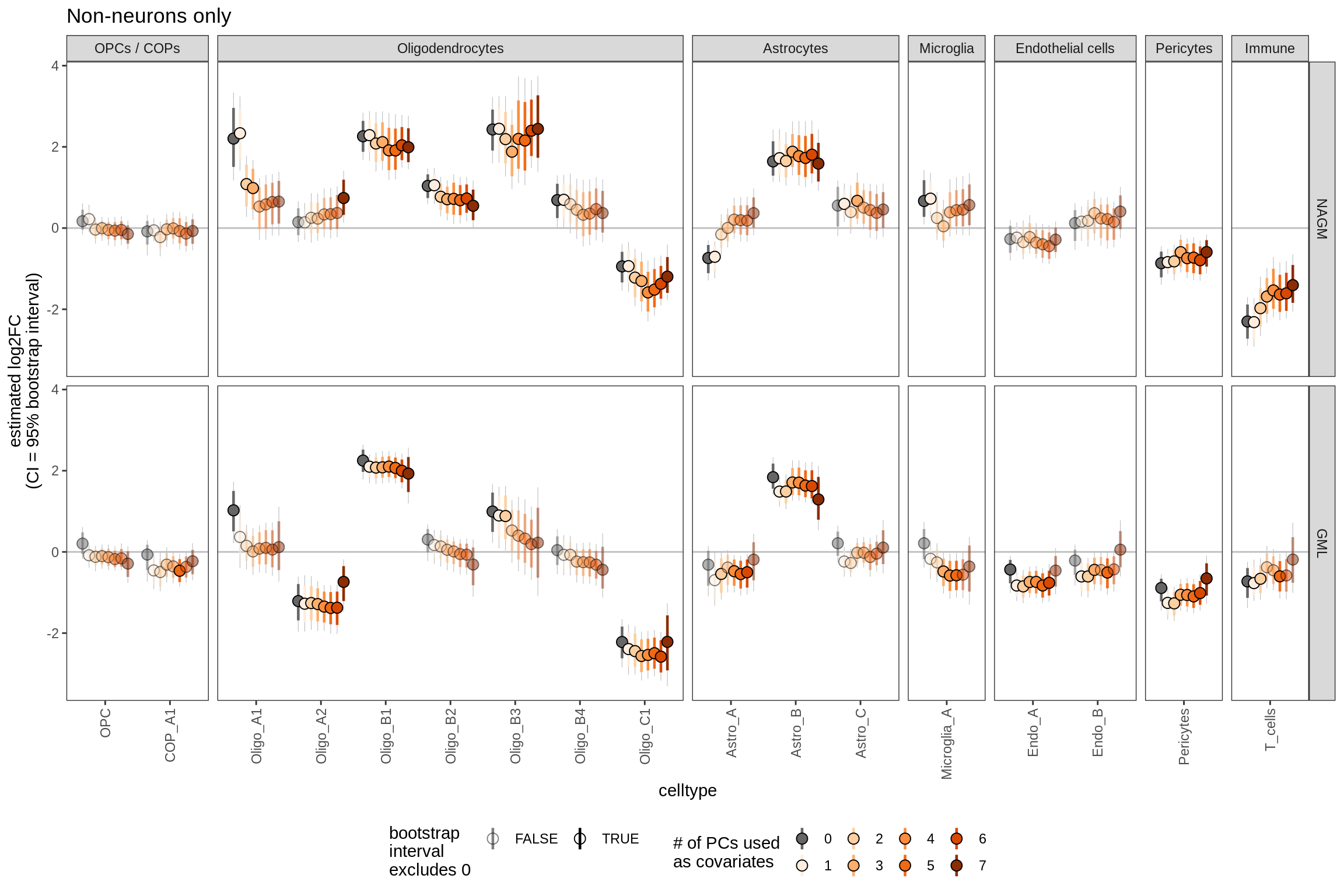
Sx
GM oligodendroglia proportions barplot
include_graphics("figure/ms09_ancombc_mixed.Rmd/plot_sample_splits_bars_oligos-2.png", error = FALSE)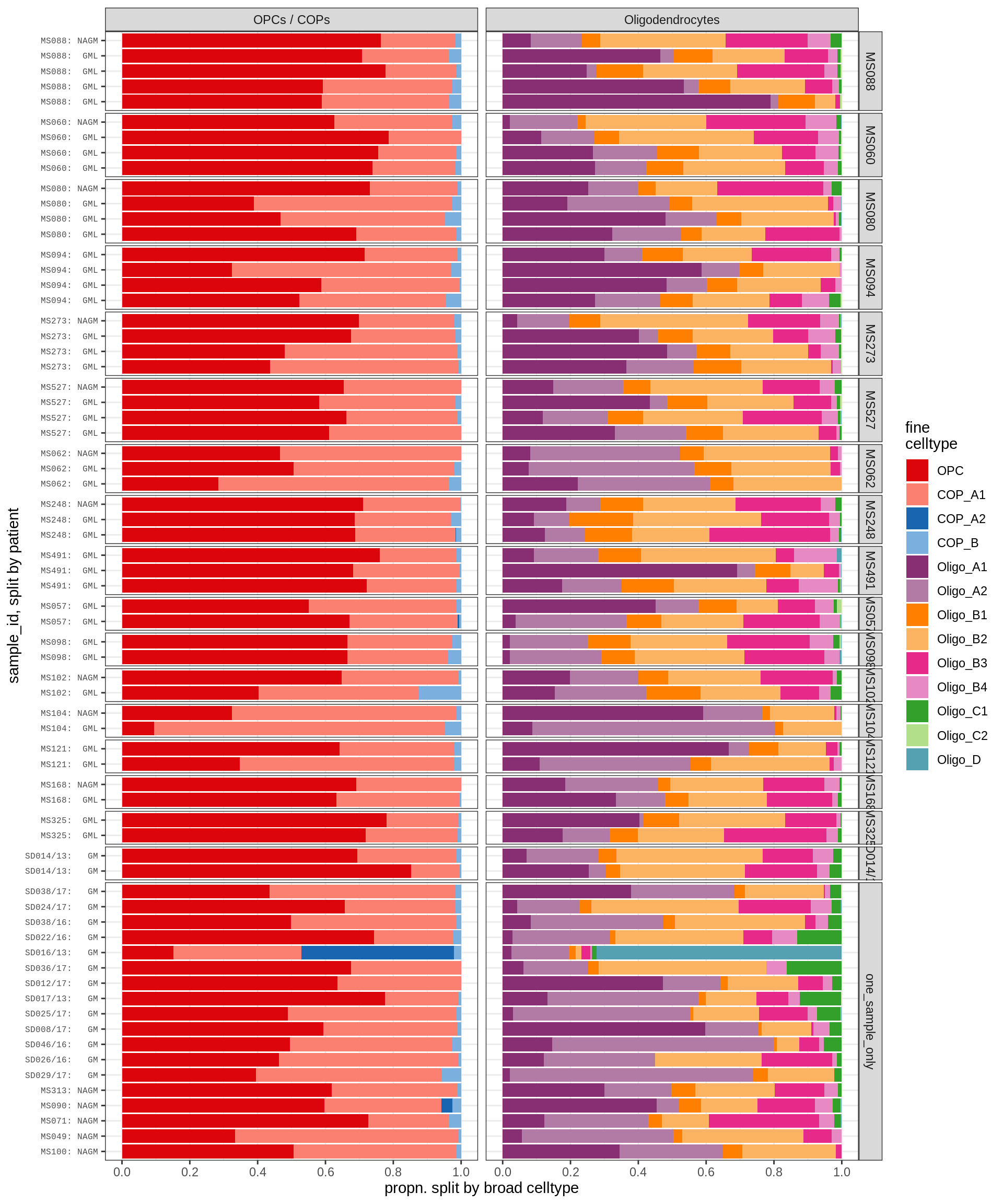
Sx
Illustration of random effects model
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/fig_random_effects_example-3.png", error = FALSE)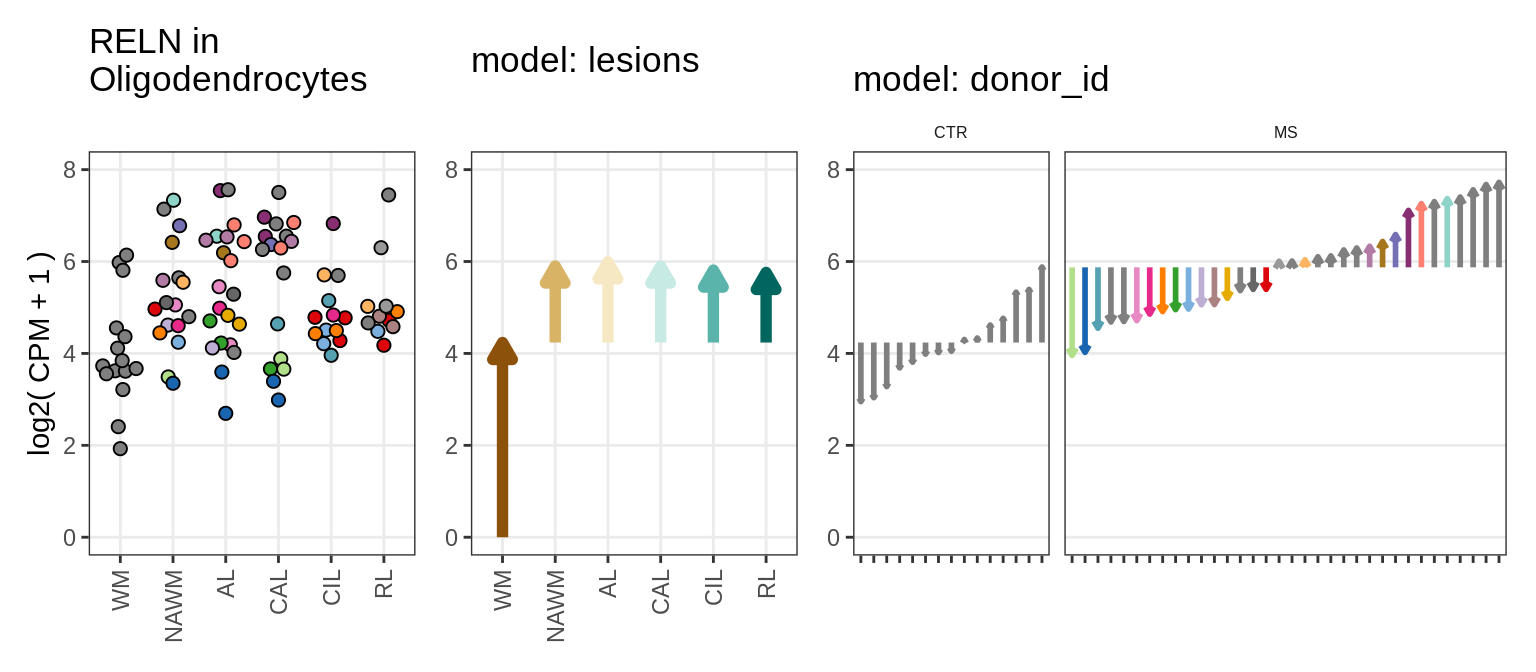
| Version | Author | Date |
|---|---|---|
| 8bc5188 | wmacnair | 2022-01-27 |
Sx
Disease effect vs donor effect
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/fig_muscat_vs_sd-1.png", error = FALSE)
Sx
MOFA factors
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/fig_mofa_factors_diagnosis-1.png", error = FALSE)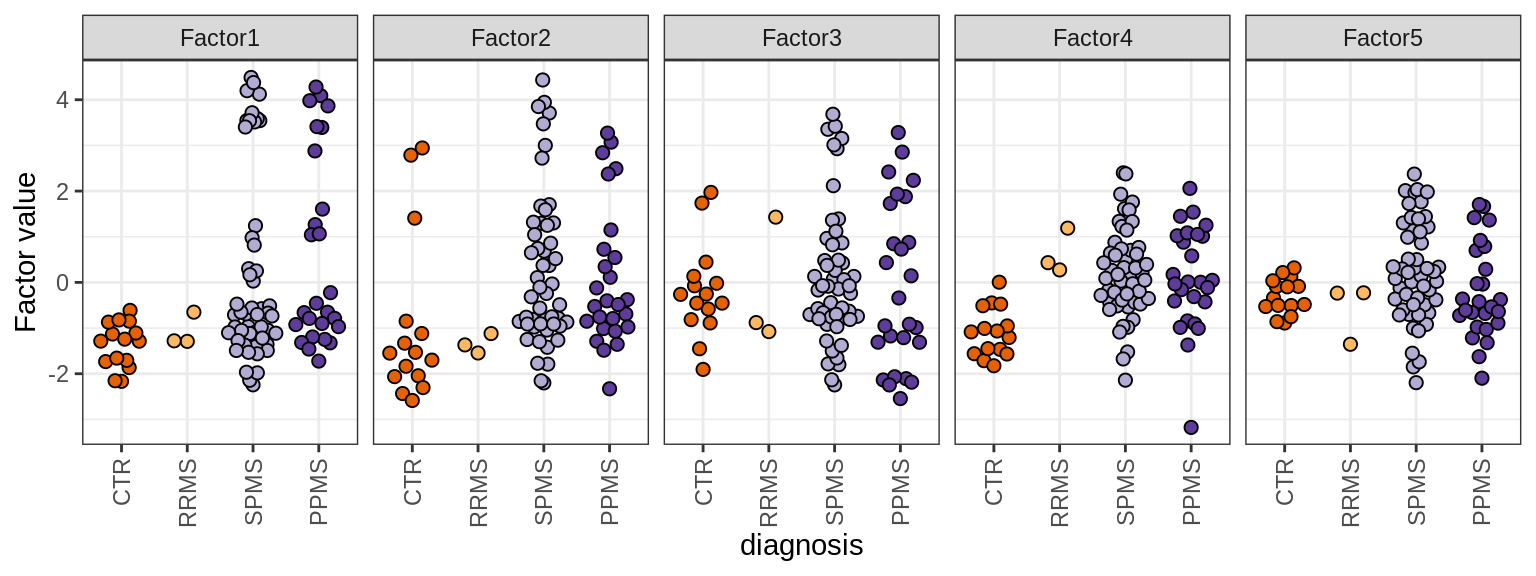
| Version | Author | Date |
|---|---|---|
| 8bc5188 | wmacnair | 2022-01-27 |
Sx
Manhattan plot of MAGMA differentially expressed genes
include_graphics("figure/gwas_figures/manhattan.png", error = FALSE)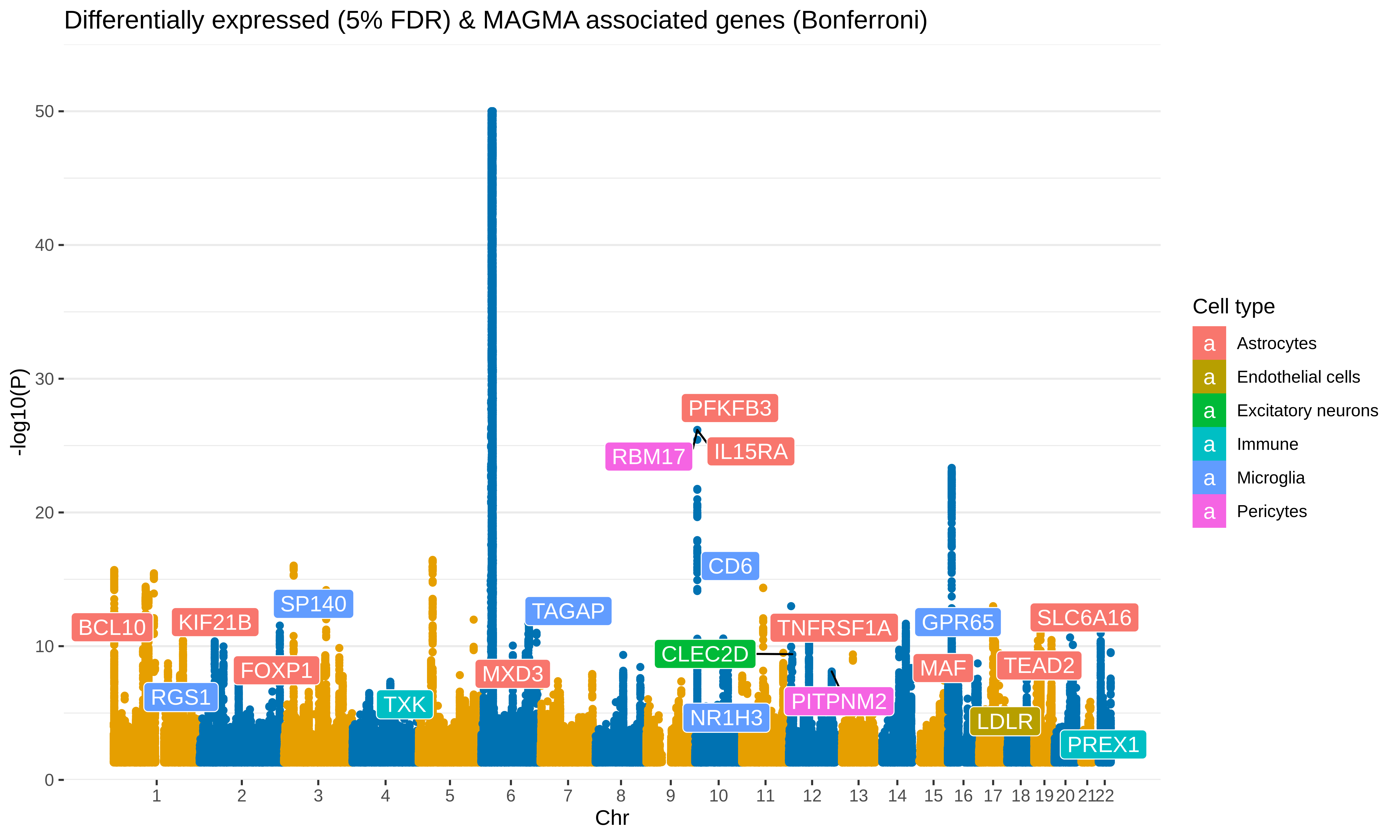
| Version | Author | Date |
|---|---|---|
| b1e52a7 | wmacnair | 2022-01-06 |
Sx
Example of a coloc gene that is differentially expressed
include_graphics("figure/gwas_figures/coloc_example_gene_NR1H3_microglia.png", error = FALSE)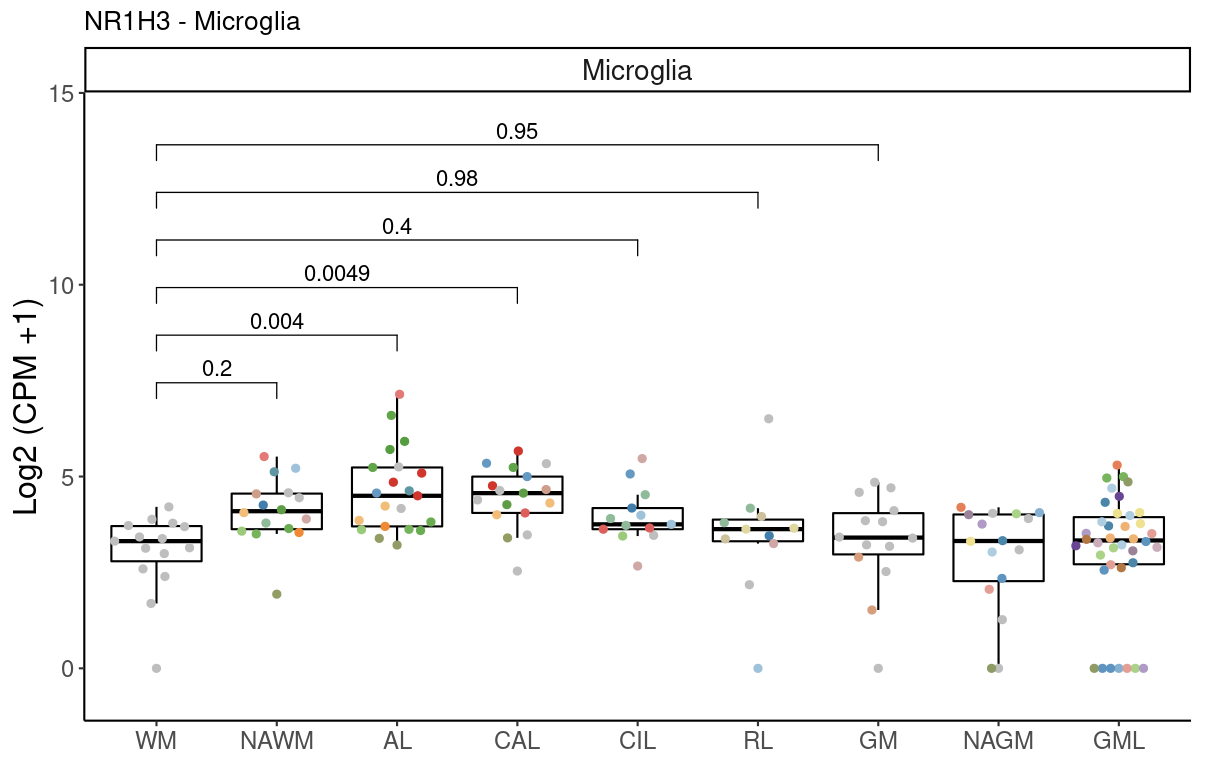
| Version | Author | Date |
|---|---|---|
| b1e52a7 | wmacnair | 2022-01-06 |
Sx
Expression heatmap of WM genes, ordered by lesion type.
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/fig_overview_expression-4.png", error = FALSE)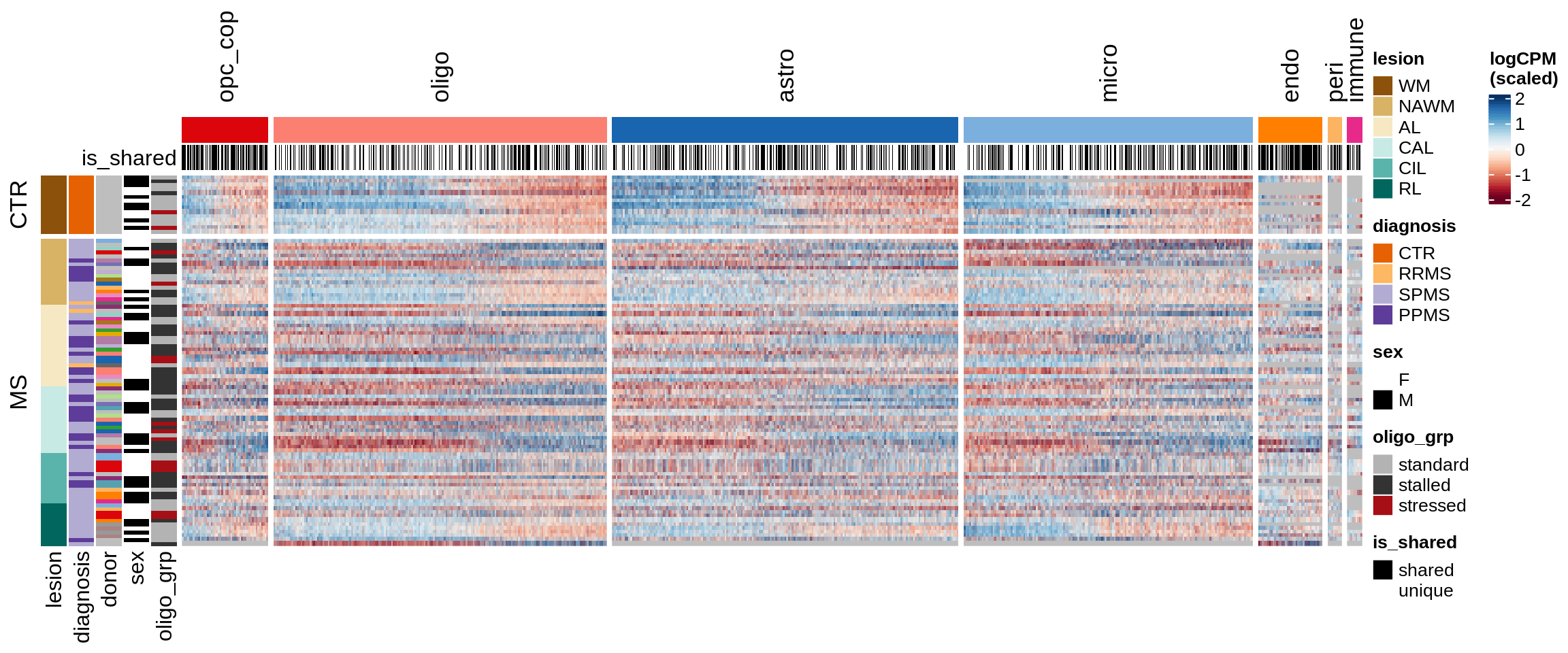
| Version | Author | Date |
|---|---|---|
| 8bc5188 | wmacnair | 2022-01-27 |
Sx
Patient stratification
include_graphics("figure/ms15_mofa_sample_wm_final_meta_bigger.Rmd/fig_f1_vs_f2-3.png", error = FALSE)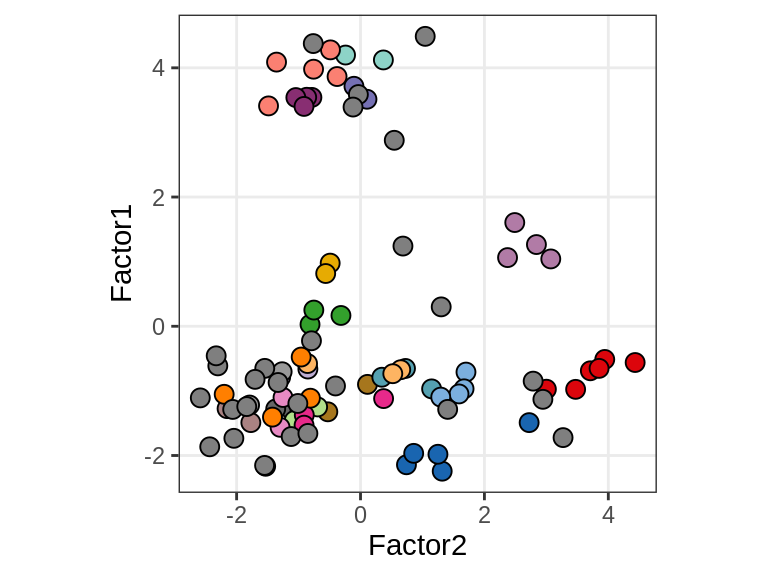
| Version | Author | Date |
|---|---|---|
| 8bc5188 | wmacnair | 2022-01-27 |
End
devtools::session_info()- Session info ---------------------------------------------------------------
setting value
version R version 4.0.5 (2021-03-31)
os CentOS Linux 7 (Core)
system x86_64, linux-gnu
ui X11
language (EN)
collate en_US.UTF-8
ctype C
tz Europe/Zurich
date 2022-01-28
- Packages -------------------------------------------------------------------
package * version date lib source
assertthat * 0.2.1 2019-03-21 [2] CRAN (R 4.0.0)
BiocManager 1.30.16 2021-06-15 [1] CRAN (R 4.0.3)
BiocStyle * 2.18.1 2020-11-24 [1] Bioconductor
bslib 0.3.1 2021-10-06 [2] CRAN (R 4.0.5)
cachem 1.0.6 2021-08-19 [1] CRAN (R 4.0.5)
callr 3.7.0 2021-04-20 [2] CRAN (R 4.0.3)
cellranger 1.1.0 2016-07-27 [2] CRAN (R 4.0.0)
circlize * 0.4.13 2021-06-09 [1] CRAN (R 4.0.3)
cli 3.0.1 2021-07-17 [1] CRAN (R 4.0.3)
codetools 0.2-18 2020-11-04 [2] CRAN (R 4.0.3)
colorout * 1.2-2 2021-04-15 [1] Github (jalvesaq/colorout@79931fd)
colorspace 2.0-2 2021-06-24 [1] CRAN (R 4.0.3)
crayon 1.4.1 2021-02-08 [2] CRAN (R 4.0.3)
data.table * 1.14.2 2021-09-27 [2] CRAN (R 4.0.5)
DBI 1.1.1 2021-01-15 [2] CRAN (R 4.0.3)
desc 1.4.0 2021-09-28 [1] CRAN (R 4.0.5)
devtools 2.4.2 2021-06-07 [1] CRAN (R 4.0.3)
digest 0.6.28 2021-09-23 [2] CRAN (R 4.0.5)
dplyr 1.0.7 2021-06-18 [2] CRAN (R 4.0.3)
ellipsis 0.3.2 2021-04-29 [2] CRAN (R 4.0.3)
evaluate 0.14 2019-05-28 [2] CRAN (R 4.0.0)
fansi 0.5.0 2021-05-25 [2] CRAN (R 4.0.3)
fastmap 1.1.0 2021-01-25 [2] CRAN (R 4.0.3)
forcats * 0.5.1 2021-01-27 [2] CRAN (R 4.0.3)
fs 1.5.0 2020-07-31 [2] CRAN (R 4.0.2)
generics 0.1.1 2021-10-25 [2] CRAN (R 4.0.5)
ggplot2 * 3.3.5 2021-06-25 [1] CRAN (R 4.0.3)
git2r 0.28.0 2021-01-10 [1] CRAN (R 4.0.3)
GlobalOptions 0.1.2 2020-06-10 [1] CRAN (R 4.0.3)
glue 1.4.2 2020-08-27 [2] CRAN (R 4.0.3)
gridExtra 2.3 2017-09-09 [2] CRAN (R 4.0.0)
gtable 0.3.0 2019-03-25 [2] CRAN (R 4.0.0)
highr 0.9 2021-04-16 [2] CRAN (R 4.0.3)
htmltools 0.5.2 2021-08-25 [2] CRAN (R 4.0.5)
httpuv 1.6.3 2021-09-09 [2] CRAN (R 4.0.5)
jquerylib 0.1.4 2021-04-26 [2] CRAN (R 4.0.3)
jsonlite 1.7.2 2020-12-09 [2] CRAN (R 4.0.3)
knitr * 1.36 2021-09-29 [1] CRAN (R 4.0.5)
later 1.3.0 2021-08-18 [2] CRAN (R 4.0.5)
lifecycle 1.0.1 2021-09-24 [2] CRAN (R 4.0.5)
magrittr * 2.0.1 2020-11-17 [1] CRAN (R 4.0.3)
memoise 2.0.0 2021-01-26 [1] CRAN (R 4.0.3)
munsell 0.5.0 2018-06-12 [2] CRAN (R 4.0.0)
pillar 1.6.4 2021-10-18 [1] CRAN (R 4.0.5)
pkgbuild 1.2.0 2020-12-15 [1] CRAN (R 4.0.3)
pkgconfig 2.0.3 2019-09-22 [2] CRAN (R 4.0.0)
pkgload 1.2.3 2021-10-13 [2] CRAN (R 4.0.5)
prettyunits 1.1.1 2020-01-24 [2] CRAN (R 4.0.0)
processx 3.5.2 2021-04-30 [2] CRAN (R 4.0.3)
promises 1.2.0.1 2021-02-11 [2] CRAN (R 4.0.3)
ps 1.6.0 2021-02-28 [2] CRAN (R 4.0.3)
purrr 0.3.4 2020-04-17 [2] CRAN (R 4.0.0)
R6 2.5.1 2021-08-19 [2] CRAN (R 4.0.5)
RColorBrewer * 1.1-2 2014-12-07 [2] CRAN (R 4.0.0)
Rcpp 1.0.7 2021-07-07 [1] CRAN (R 4.0.3)
readxl * 1.3.1 2019-03-13 [2] CRAN (R 4.0.0)
remotes 2.4.1 2021-09-29 [1] CRAN (R 4.0.5)
rlang 0.4.12 2021-10-18 [2] CRAN (R 4.0.5)
rmarkdown 2.11 2021-09-14 [1] CRAN (R 4.0.5)
rprojroot 2.0.2 2020-11-15 [2] CRAN (R 4.0.3)
sass 0.4.0 2021-05-12 [2] CRAN (R 4.0.3)
scales * 1.1.1 2020-05-11 [2] CRAN (R 4.0.0)
sessioninfo 1.1.1 2018-11-05 [1] CRAN (R 4.0.3)
shape 1.4.6 2021-05-19 [1] CRAN (R 4.0.1)
stringi 1.7.4 2021-08-25 [1] CRAN (R 4.0.5)
stringr * 1.4.0 2019-02-10 [2] CRAN (R 4.0.0)
testthat 3.1.0 2021-10-04 [2] CRAN (R 4.0.5)
tibble 3.1.5 2021-09-30 [1] CRAN (R 4.0.5)
tidyselect 1.1.1 2021-04-30 [2] CRAN (R 4.0.3)
usethis 2.1.2 2021-10-25 [1] CRAN (R 4.0.5)
utf8 1.2.2 2021-07-24 [1] CRAN (R 4.0.3)
vctrs 0.3.8 2021-04-29 [2] CRAN (R 4.0.3)
viridis * 0.6.2 2021-10-13 [1] CRAN (R 4.0.5)
viridisLite * 0.4.0 2021-04-13 [1] CRAN (R 4.0.1)
whisker 0.4 2019-08-28 [1] CRAN (R 4.0.3)
withr 2.4.2 2021-04-18 [2] CRAN (R 4.0.3)
workflowr * 1.6.2 2020-04-30 [1] CRAN (R 4.0.3)
xfun 0.27 2021-10-18 [1] CRAN (R 4.0.5)
yaml 2.2.1 2020-02-01 [2] CRAN (R 4.0.3)
[1] /pstore/home/macnairw/lib/conda_r3.12
[2] /pstore/home/macnairw/.conda/envs/r_4.0.3/lib/R/library
sessionInfo()R version 4.0.5 (2021-03-31)
Platform: x86_64-conda-linux-gnu (64-bit)
Running under: CentOS Linux 7 (Core)
Matrix products: default
BLAS/LAPACK: /pstore/home/macnairw/.conda/envs/r_4.0.3/lib/libopenblasp-r0.3.12.so
locale:
[1] LC_CTYPE=C LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] knitr_1.36 readxl_1.3.1 forcats_0.5.1 ggplot2_3.3.5
[5] scales_1.1.1 viridis_0.6.2 viridisLite_0.4.0 assertthat_0.2.1
[9] stringr_1.4.0 data.table_1.14.2 magrittr_2.0.1 circlize_0.4.13
[13] RColorBrewer_1.1-2 BiocStyle_2.18.1 colorout_1.2-2 workflowr_1.6.2
loaded via a namespace (and not attached):
[1] Rcpp_1.0.7 prettyunits_1.1.1 ps_1.6.0
[4] rprojroot_2.0.2 digest_0.6.28 utf8_1.2.2
[7] R6_2.5.1 cellranger_1.1.0 evaluate_0.14
[10] highr_0.9 pillar_1.6.4 GlobalOptions_0.1.2
[13] rlang_0.4.12 callr_3.7.0 whisker_0.4
[16] jquerylib_0.1.4 rmarkdown_2.11 desc_1.4.0
[19] devtools_2.4.2 munsell_0.5.0 compiler_4.0.5
[22] httpuv_1.6.3 xfun_0.27 pkgconfig_2.0.3
[25] pkgbuild_1.2.0 shape_1.4.6 htmltools_0.5.2
[28] tidyselect_1.1.1 tibble_3.1.5 gridExtra_2.3
[31] codetools_0.2-18 fansi_0.5.0 crayon_1.4.1
[34] dplyr_1.0.7 withr_2.4.2 later_1.3.0
[37] grid_4.0.5 jsonlite_1.7.2 gtable_0.3.0
[40] lifecycle_1.0.1 DBI_1.1.1 git2r_0.28.0
[43] cli_3.0.1 stringi_1.7.4 cachem_1.0.6
[46] remotes_2.4.1 fs_1.5.0 promises_1.2.0.1
[49] testthat_3.1.0 bslib_0.3.1 ellipsis_0.3.2
[52] generics_0.1.1 vctrs_0.3.8 tools_4.0.5
[55] glue_1.4.2 purrr_0.3.4 pkgload_1.2.3
[58] processx_3.5.2 fastmap_1.1.0 yaml_2.2.1
[61] colorspace_2.0-2 BiocManager_1.30.16 sessioninfo_1.1.1
[64] memoise_2.0.0 usethis_2.1.2 sass_0.4.0